Small Interfering RNA Delivery Into Primary Human Natural Killer Cells for Functional Gene Analyses
Pouria Momayyezi, Pouria Momayyezi, Karl-Johan Malmberg, Karl-Johan Malmberg, Quirin Hammer, Quirin Hammer
Abstract
Studying gene functions in human natural killer (NK) cells is key for advancing the understanding of NK cell biology and holds promise to pave the way for improving NK cell therapies against cancer. However, NK cells are challenging to manipulate, and investigation of gene functions in NK cells is hampered by variable delivery efficiencies and impaired viability upon electroporation, lipofection, or viral transduction. Here, we report a simple workflow for delivery of commercially available small interfering RNA molecules into primary human NK cells to enable functional gene analyses. © 2022 The Authors. Current Protocols published by Wiley Periodicals LLC.
Basic Protocol 1 : Enrichment of natural killer cells from human peripheral blood mononuclear cells
Basic Protocol 2 : Preparation of small interfering RNA
Basic Protocol 3 : Delivery of small interfering RNA into natural killer cells
Support Protocol 1 : Isolation of human peripheral blood mononuclear cells from buffy coats
Support Protocol 2 : Thawing and recovery of cryopreserved peripheral blood mononuclear cells
Support Protocol 3 : Evaluation of natural killer cell purity following magnetic enrichment
Support Protocol 4 : Exemplary assessment of knockdown efficiency
INTRODUCTION
As part of the innate immune system, natural killer (NK) cells represent one of the first lines of host defense against virus infections and developing tumors (Bjorkstrom, Strunz, & Ljunggren, 2022; Wolf, Kissiov, & Raulet, 2022). The antitumor activity of NK cells is being explored clinically, for example by stimulating endogenous NK cells of the patient or by adoptive transfer of NK cells as therapeutic product (Chiossone & Vivier, 2022). Thus, a better understanding of gene functions in NK cells will shed light on fundamental NK cell biology and holds potential for improving the translational value of NK cells within cancer therapy. RNA interference–mediated transient silencing of gene expression by transfer of small interfering RNA (siRNA) molecules, which target messenger RNA sequences of the gene of interest, is a powerful tool for functional gene analysis in a loss-of-function manner.
Here, we provide a detailed protocol for the delivery of commercially available siRNA into primary human NK cells. A critical aspect of the transfer of siRNA molecules into cells is that the use of chemically modified siRNA enables passive uptake and does not involve procedures that diminish viability such as electroporation. Our passive uptake approach results in high delivery efficiency while sustaining NK cell viability and function (Palacios et al., 2022), thereby extending the molecular toolkit with a valuable technique to study NK cell biology.
We first describe a protocol for the enrichment of NK cells from human peripheral blood mononuclear cells (PBMCs) by magnetic depletion of non-NK cell populations (Basic Protocol 1). We then detail the preparation of Accell siRNA reagents (Basic Protocol 2) and describe the procedure for delivering Accell siRNA into NK cells (Basic Protocol 3).
Furthermore, we include information for isolating human PBMCs from leukocyte concentrates (“buffy coats;” Support Protocol 1) and for thawing and recovering PBMCs after cryopreservation (Support Protocol 2). We also include a protocol for flow cytometric evaluation of cell purity after NK cell isolation (Support Protocol 3) and describe procedures to assess knockdown efficiency after an exemplary treatment with siRNA reagents that target the expression of the beta-2-microglobulin (B2M) gene (Support Protocol 4).
CAUTION : When working with human material, safety precautions must be followed, such as wearing appropriate personal protective equipment, using disinfectant, and working at the correct biosafety level. Consult with your institution's biological safety department to ensure that all proper precautions are taken.
NOTE : All protocols involving humans and human samples must first be reviewed and approved by an institutional review board (IRB) or must follow local guidelines for the use of human samples. All participants must provide informed consent.
NOTE : These protocols assume familiarity with standard tissue culture methods and the availability of equipment for such procedures including a biosafety cabinet, centrifuge, hemocytometer (manual or automated), water bath, ice machine, –80°C freezer, liquid nitrogen storage, tissue culture incubator, standard laboratory plastic ware, and flow cytometry and analysis software.
Basic Protocol 1: ENRICHMENT OF NATURAL KILLER CELLS FROM HUMAN PERIPHERAL BLOOD MONONUCLEAR CELLS
This protocol provides instructions for the enrichment of CD3– CD14– CD19– CD56+ NK cells from human PBMCs by negative selection through magnetic labeling and depletion of non-NK cell populations. Alternatively, NK cells or specific NK cell subsets can be purified by other ways such as fluorescence-activated cell sorting (FACS).
Materials
- Human PBMCs (e.g., obtained by density gradient centrifugation and used directly [see Support Protocol 1] or cryopreserved and thawed [see Support Protocol 2])
- Trypan blue solution (e.g., Gibco, cat. no. 15250061)
- MACS buffer (see recipe)
- NK Cell Isolation Kit, human (e.g., Miltenyi Biotec, cat. no. 130-092-657) containing:
- NK Cell Biotin-Antibody Cocktail
- NK Cell MicroBead Cocktail
- Phosphate-buffered saline (PBS), pH 7.4 (e.g., Gibco, cat. no. 10010023)
- Hemocytometer
- 15-ml conical tubes (e.g., Corning, cat. no. 352096)
- Centrifuge
- QuadroMACS Separator (e.g., Miltenyi Biotec, cat. no. 130-090-976)
- MACS MultiStand (e.g., Miltenyi Biotec, cat. no. 130-042-303)
- LS columns (e.g., Miltenyi Biotec, cat. no. 130-042-401)
- Insulated ice container for wet ice (e.g., NeoLab cat. no. 2-4994)
Prepare PBMCs
1.Obtain human PBMCs by method of choice.
2.Determine number of viable PBMCs by trypan blue exclusion in a hemocytometer.
3.Transfer 50 × 106 viable PBMCs to a 15-ml conical tube.
4.Centrifuge 10 min at 300 × g , 4°C.
5.Remove supernatant completely, and resuspend cell pellet in 200 µl cold MACS buffer.
Label PBMCs
6.Add 50 µl cold NK Cell Biotin-Antibody Cocktail and mix carefully.
7.Incubate for 5 min at 4°C.
8.Without centrifugation, add 150 µl cold MACS buffer and 100 µl cold NK Cell MicroBead Cocktail and mix carefully.
9.Incubate for 10 min at 4°C.
10.During incubation, prepare for magnetic enrichment:
- a.Place a QuadroMACS separator on the MACS MultiStand.
- b.Insert an LS column into the separator.
- c.Place a bucket with wet ice under the separator.
- d.Rinse column with 3 ml cold MACS buffer and discard flow-through.
- e.
Prepare a new 15-ml conical tube for collection of the enriched NK cells by coating the walls of the tube with cold MACS buffer. Place coated tube below the column into the ice bucket.
Coating the wall of the tube reduces cell loss. Add ∼2 ml cold MACS buffer to the conical tube, and rotate carefully until all areas are covered with liquid.
Deplete non-NK cells
11.At the end of the incubation period, transfer suspension of magnetically labeled PBMCs to the column.
12.Let suspension run through the column by gravity without applying force, and collect flow-through, which contains the non-labeled NK cells.
13.Rinse column with 3 ml cold MACS buffer.
14.After rinsing is completed, carefully remove 15-ml conical tube containing the flow-through, and centrifuge 10 min at 300 × g , 4°C.
15.Remove supernatant and resuspend enriched NK cells in 1 ml warm PBS.
16.Determine number of obtained viable NK cells by trypan blue exclusion in a hemocytometer.
Basic Protocol 2: PREPARATION OF SMALL INTERFERING RNA
This protocol provides information for reconstituting lyophilized Accell siRNA. Successful reconstitution is critical for downstream applications. Reconstitution of siRNA can be performed immediately before siRNA treatment of enriched NK cells. Alternatively, reconstitution can be performed beforehand, and the reconstituted siRNA can be stored at –20°C until treatment of NK cells.
Materials
- 5× siRNA buffer (e.g., Horizon Discovery, cat. no. B-002000-UB-100)
- RNase-free water (e.g., Horizon Discovery, cat. no. B-003000-WB-100)
- Accell siRNA of choice (e.g., Horizon Discovery)
- Microcentrifuge
- Laboratory shaker
- Spectrophotometer
- RNase-free reaction tubes (e.g., Sarstedt, cat. no. 72.706.200)
NOTE : This protocol assumes familiarity with standard molecular biology techniques and the availability of equipment for such procedures including a centrifuge and an orbital shaker suitable for 1.5-ml reaction tubes, RNase-free environment, and PCR-grade pipet tips.
1.Bring 5× siRNA buffer and RNase-free water to room temperature.
2.Dilute 5× siRNA buffer with RNase-free water to 1× siRNA buffer.
3.Briefly centrifuge vial containing lyophilized siRNA at 14,000 × g , room temperature.
4.Add 50 µl of 1× siRNA buffer to 5 nmol lyophilized siRNA to obtain a stock concentration of 100 µM.
5.Mix by carefully pipetting up and down five times without introducing air bubbles into the solution.
6.Incubate solution for 30 min at room temperature while shaking at 50 rpm.
7.Briefly centrifuge vial containing the siRNA solution at 1000 × g , room temperature, to collect any drops that may have formed during reconstitution.
8.Verify reconstitution by determining the concentration of siRNA in solution (e.g., using a spectrophotometer).
9.Use reconstituted siRNA immediately for delivery into NK cells (Basic Protocol 3), or aliquot into RNase-free reaction tubes to minimize freeze-thaw cycles, storing at −20°C until use.
Basic Protocol 3: DELIVERY OF SMALL INTERFERING RNA INTO NATURAL KILLER CELLS
This protocol details the delivery of Accell siRNA into enriched human NK cells. Uptake of Accell siRNA is a passive process and does not require electroporation, transfection reagents, or viral vectors. Serum-free Accell siRNA delivery medium strongly enhances siRNA uptake and, when supplemented with low doses of recombinant human IL-15 as a prosurvival factor, does not compromise the viability of human NK cells throughout the treatment period.
Materials
- PBS, pH 7.4 (e.g., Gibco, cat. no. 10010023)
- Accell siRNA delivery medium (e.g., Horizon Discovery, cat. no. B-005000-100)
- 10,000 U/ml penicillin/10,000 μg/ml streptomycin (e.g., Gibco, cat. no. 15140122)
- Recombinant human IL-15, premium grade (e.g., Miltenyi Biotec, cat. no. 130-095-762)
- Enriched NK cells (see Basic Protocol 1)
- 37°C water bath
- 15-ml conical tubes (e.g., Corning, cat. no. 352096)
- Centrifuge
- 96-well U-bottom tissue culture plates (e.g., Corning, cat. no. 353077)
- 37°C tissue culture incubator, 5% CO2, 95% humidity
1.Bring PBS and Accell siRNA delivery medium to 37°C using a water bath.
2.Supplement warm Accell siRNA delivery medium with 100 U/ml penicillin, 100 μg/ml streptomycin, and 1 ng/ml recombinant human IL-15.
3.Transfer 1 × 106 enriched NK cells into a 15-ml conical tube, and add 5 ml warm PBS. Centrifuge 10 min at 300 × g , room temperature.
4.Remove supernatant from the cell pellet completely.
5.Resuspend NK cells in 1 ml warm Accell siRNA delivery medium containing all desired supplements.
6.Dispense 100 μl cell suspension (containing 1 × 105 NK cells) into the wells of a 96-well U-bottom plate.
7.Add 1 μl Accell siRNA directly into the wells containing the cell suspension and mix.
8.Incubate plate in a tissue culture incubator at 37°C with 5% CO2 and 95% humidity for 48 to 96 hr.
Support Protocol 1: ISOLATION OF HUMAN PERIPHERAL BLOOD MONONUCLEAR CELLS FROM BUFFY COATS
This protocol describes the isolation of human PBMCs from leukocyte concentrates (“buffy coats”) through density gradient centrifugation. At step 13, the obtained PBMCs can be used directly for downstream applications (e.g., Basic Protocol 1). Alternatively, PBMCs can be cryopreserved for later use.
Please note that several other methods are available for isolation of PBMCs.
Materials
- Buffy coat (obtained from volunteer blood bank donors)
- PBS, pH 7.4 (e.g., Gibco, cat. no. 10010023)
- Ficoll-Paque PLUS (e.g., Cytiva, cat. no. 17144002)
- Trypan blue solution (e.g., Gibco, cat. no. 15250061)
- Cryostorage medium (see recipe)
- 175-cm2 flask (e.g., Corning, cat. no. 431080)
- 50-ml conical tube (e.g. Corning, cat. no. 352098)
- Centrifuge
- Hemocytometer
- Cryotubes (e.g., Sarstedt, cat. no. 72.377.002)
- Alcohol-resistant, low-temperature-stable marker or label maker
- Alcohol-free cell freezing container (e.g., Corning, cat. no. 432004)
- Liquid nitrogen storage container
Isolate PBMCs
1.Transfer entire buffy coat sample to a 175-cm2 flask.
2.Add PBS to the sample, and dilute to a final volume of 140 ml.
3.Add 15 ml Ficoll-Paque PLUS to four 50-ml conical tubes.
4.Carefully layer 35 ml diluted buffy coat sample onto the Ficoll-Paque PLUS in each tube.
5.Centrifuge 20 min at 800 × g , room temperature, with deceleration turned off.
6.Remove and discard the top 20 to 25 ml plasma layer.
7.Carefully collect PBMC layer at the interface of the plasma layer and Ficoll-Paque, and transfer to a new 50-ml conical tube.
8.Add PBS to a final volume of 50 ml, and centrifuge 10 min at 300 × g , room temperature.
9.Remove supernatant.
10.Resuspend cell pellet in 50 ml PBS, and centrifuge 15 min at 200 × g , room temperature.
11.Remove supernatant.
12.Resuspend cell pellet in 50 ml PBS.
13.Determine number of viable PBMCs by trypan blue exclusion in a hemocytometer.
Optional: Cryopreserve PBMCs
14.Prepare cryostorage medium.
15.Prepare appropriate number of cryotubes, and label with an alcohol-resistant, low-temperature-stable marker or printed label.
16.Centrifuge PBMC suspension 10 min at 300 × g , room temperature.
17.Remove supernatant completely.
18.Resuspend cell pellet in an appropriate volume of cryostorage medium and mix carefully.
19.Dispense 1 ml cell suspension into the prepared cryotubes.
20.Immediately transfer cryotubes to an alcohol-free cell freezing container, and place container at −80°C.
21.After 3 to 16 hr, transfer cryovials to liquid nitrogen storage (less than −140°C) for long-term storage.
Support Protocol 2: THAWING AND RECOVERY OF CRYOPRESERVED PERIPHERAL BLOOD MONONUCLEAR CELLS
This protocol describes a procedure for thawing and recovering previously cryopreserved PBMCs. At the final step of the protocol, the obtained PBMCs can be used directly for downstream applications such as enrichment of NK cells (Basic Protocol 1).
Materials
- Culture medium (see recipe)
- 250 U/μl benzonase nuclease (e.g., Merck, cat. no. E1014-5KU)
- Cryopreserved PBMCs (see Support Protocol 1)
- Trypan blue solution (e.g., Gibco, cat. no. 15250061)
- 37°C water bath
- 50-ml conical tube (e.g., Corning, cat. no. 352098)
- Centrifuge
- 70-µm cell strainer (e.g., Corning, cat. no. 352350)
- Hemocytometer
- 6-well tissue culture plate (e.g., Corning, cat. no. 353046)
- 37°C tissue culture incubator, 5% CO2, 95% humidity
Thaw PBMCs
1.Prepare culture medium, and bring to 37°C using a water bath.
2.For each sample, transfer 20 ml culture medium to a 50-ml conical tube.
3.Add 2 μl benzonase nuclease to the culture medium.
4.Collect cryopreserved PBMC samples from liquid nitrogen storage.
5.Immediately place sample into a 37°C water bath.
6.Quickly thaw cells until only a small piece of ice remains visible.
7.Transfer thawed cells dropwise to the tube with warm culture medium containing benzonase.
8.Add 1 ml warm culture medium containing benzonase to the cryovial.
9.Pipet up and down once to thaw remaining ice, and transfer thawed cells to the tube with culture medium containing benzonase.
10.Centrifuge 10 min at 300 × g , room temperature.
11.Remove supernatant.
12.Resuspend cell pellet in 20 ml warm culture medium without benzonase, and strain cell suspension through a 70-µm cell strainer.
13.Centrifuge 10 min at 300 × g , room temperature.
14.Remove supernatant.
15.Resuspend cell pellet in 20 ml warm culture medium.
Recover thawed PBMCs
16.Determine number of viable PBMCs by trypan blue exclusion in a hemocytometer.
17.Adjust cell density to 5 × 106 cells/ml, and dispense 5 ml to wells of a 6-well tissue culture plate.
18.Incubate plate in a tissue culture incubator at 37°C with 5% CO2 and 95% humidity for 2 to 18 hr.
19.After incubation, gather cells from plate, and transfer to a 50-ml conical tube.
20.Determine number of viable PBMCs by trypan blue exclusion in a hemocytometer.
Support Protocol 3: EVALUATION OF NATURAL KILLER CELL PURITY FOLLOWING MAGNETIC ENRICHMENT
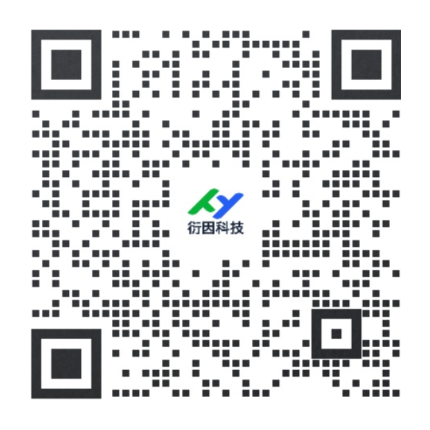
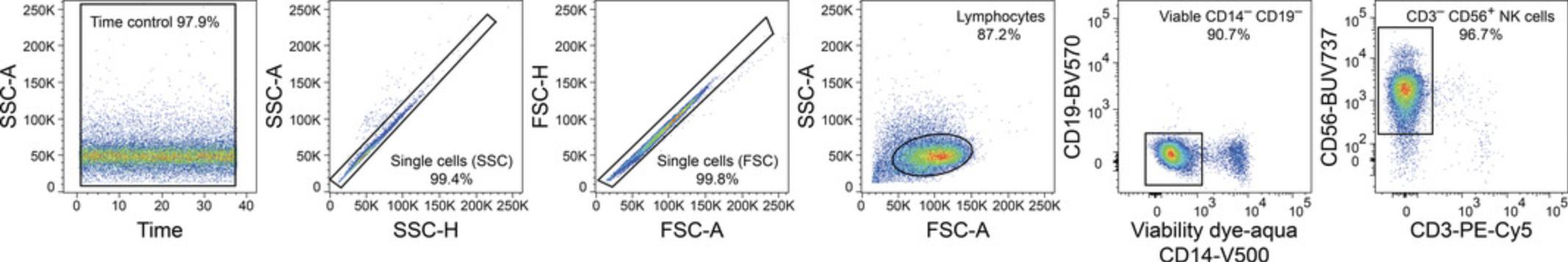
This protocol describes a flow cytometric evaluation of viable CD14– CD19– CD3– CD56+ NK cells following magnetic enrichment (Basic Protocol 1) to determine the purity of the obtained population.
Materials
- Viability dye (e.g., LIVE/DEAD Fixable Aqua Dead Cell Stain Kit; Invitrogen, cat. no. L34957)
- Fluorochrome-conjugated antibodies:
- V500-conjguated anti-human CD14, clone M5E2 (e.g., BD Biosciences, cat. no. 561391; RRID AB_10611856)
- BV570-conjugated anti-human CD19, clone HIB19 (e.g., BioLegend, cat. no. 302236; RRID AB_2563606)
- PE-Cy5-conjguated anti-human CD3, clone UCHT1 (e.g., Beckman Coulter, cat. no. A07749; RRID AB_2827419)
- BUV737-conjugated anti-human CD56, clone NCAM16.2 (e.g., BD Biosciences, cat. no. 612767; RRID AB_2860005)
- PBS (e.g., Gibco, cat. no. 10-010-23)
- Enriched NK cells (see Basic Protocol 1)
- Fixation buffer (e.g., Cytofix; BD Biosciences, cat. no. 554655)
- FACS buffer (see recipe)
- 96-well conical microwell plate with lid (e.g., Nunc, cat. nos. 249570 and 263339, respectively)
- Centrifuge
- Flow cytometer and analysis software
CAUTION : When working with fixatives that contain paraformaldehyde, proper safety precautions and waste handling steps must be followed.
1.Prepare staining solution containing viability dye and fluorochrome-conjugated antibodies at optimal dilutions in PBS.
2.Transfer 1–5 × 105 enriched NK cells to a 96-well conical microwell plate with lid.
3.Add PBS to each well to a final volume of 200 μl.
4.Centrifuge 5 min at 300 × g , room temperature.
5.Remove supernatant.
6.Add 50 μl staining solution to the cell pellet.
7.Mix well and incubate for 20 min at room temperature in the dark.
8.Wash cells twice in PBS:
-
For the first wash, add 150 μl PBS to the wells. Centrifuge 5 min at 300 ×g, room temperature, and remove supernatant.
-
For the second wash, resuspend cell pellet in 200 μl PBS. Centrifuge 5 min at 300 ×g, room temperature, and remove supernatant.
9.Resuspend cell pellet in 100 μl fixation buffer.
10.Incubate for 15 min at 4°C in the dark.
11.Wash cells twice in FACS buffer:
-
For the first wash, add 150 μl FACS buffer to the wells. Centrifuge 5 min at 500 ×g, room temperature, and remove supernatant.
-
For the second wash, resuspend cell pellet in 200 μl FACS buffer. Centrifuge 5 min at 500 ×g, room temperature, and remove supernatant.
12.Resuspend cell pellet in 100 μl FACS buffer.
13.Analyze sample by flow cytometry.
14.Use analysis software to evaluate the purity of viable CD14– CD19– CD3– CD56+ NK cells within single-lymphocyte events.
Support Protocol 4: EXEMPLARY ASSESSMENT OF KNOCKDOWN EFFICIENCY
This protocol details a procedure to assess the knockdown efficiency of primary human NK cells treated with different siRNA reagents that target transcripts of the B2M gene. For assessment of knockdown efficiency, β2m protein expression at the cell surface is analyzed by flow cytometry. In parallel, cell surface expression of human leukocyte antigen (HLA) class I protein, which is stabilized in the presence of β2m, is also determined on the siRNA-treated cells. Various other methods are available to assess knockdown efficiency, both at transcript level (e.g., reverse-transcription quantitative PCR) or protein level (e.g., immunoblot). Choosing an appropriate method for determining knockdown efficiency will be highly dependent on the nature of the gene of interest and the study design.
In this example, human PBMCs were isolated from buffy coats and cryopreserved (Support Protocol 1), thawed and recovered (Support Protocol 2), enriched for NK cells (Basic Protocol 1), and treated with siRNA (Basic Protocol 3).
Treatment with siRNA was performed with the following conditions:
- (1) control nontargeting siRNA (siRNA with minimal targeting of human genes to determine baseline responses to treatment);
- (2) B2M siRNA SMARTpool (pool of four siRNA sequences [nos. 13, 14, 15, and 16] that target the B2M gene);
- (3) B2M siRNA individual sequence 13;
- (4) B2M siRNA individual sequence 14;
- (5) B2M siRNA individual sequence 15; and
- (6) B2M siRNA individual sequence 16.
Materials
- Viability dye (e.g., LIVE/DEAD Fixable Aqua Dead Cell Stain Kit; Invitrogen, cat. no. L34957)
- Fluorochrome-conjugated antibodies:
- PE-Cy5-conjguated anti-human CD3, clone UCHT1 (e.g., Beckman Coulter, cat. no. A07749; RRID AB_2827419)
- BUV737-conjugated anti-human CD56, clone NCAM16.2 (e.g., BD Biosciences, cat. no. 612767; RRID AB_2860005)
- APC-conjugated isotype control, clone MOPC-21 (e.g., BioLegend, cat no. 400119; RRID AB_2888687)
- FITC-conjugated isotype control, clone MOPC-21 (e.g., BioLegend, cat. no. 400107; RRID AB_326429)
- APC-conjugated anti-human β2m, clone 2M2 (e.g., BioLegend, cat. no. 316312; RRID AB_10641281)
- FITC-conjugated anti-human HLA class I, clone W6/32 (e.g., BioLegend, cat. no. 311404; RRID AB_314873)
- PBS (e.g., Gibco, cat. no. 10-010-23)
- NK cells treated with the following siRNA (see Basic Protocol 3):
- Nontargeting control siRNA (e.g., Horizon Discovery, cat. no. D-001910-10-05)
- B2M siRNA SMARTpool (mix of four siRNA sequences; e.g., Horizon Discovery, cat. no. E-004366-00-0005)
- B2M siRNA individual 13 (e.g., Horizon Discovery, cat. no. A-004366-13-0005)
- B2M siRNA individual 14 (e.g., Horizon Discovery, cat. no. A-004366-14-0005)
- B2M siRNA individual 15 (e.g., Horizon Discovery, cat. no. A-004366-15-0005)
- B2M siRNA individual 16 (e.g., Horizon Discovery, cat. no. A-004366-16-0005)
- Fixation buffer (e.g., Cytofix; BD Biosciences, cat. no. 554655)
- FACS buffer (see recipe)
- 96-well conical microwell plate plus lid (e.g., Nunc, cat. nos. 249570 and 263339, respectively)
- Centrifuge
- Flow cytometer and analysis software
CAUTION : When working with fixatives that contain paraformaldehyde, proper safety precautions and waste handling steps must be followed.
1.Prepare staining solutions containing viability dye and fluorochrome-conjugated antibodies at optimal dilutions in PBS:
- a.Prepare a staining solution containing viability dye, anti-human CD3-PE-Cy5, anti-human CD56-BUV737, isotype control-APC, and isotype control-FITC.
- b.
Prepare a staining solution containing viability dye, anti-human CD3-PE-Cy5, anti-human CD56-BUV737, anti-human β2m-APC, and anti-human HLA class I-FITC.
Isotype controls indicate the unspecific fluorescence signal and thus serve as important controls for determining baseline signals in the evaluation of knockdown efficiency.
The manufacturers provide a recommended dilution for each antibody. However, additional titration is required to define the optimal working concentration for each reagent. The following dilutions have been identified as optimal in our laboratory: fixable viability dye, 1:100; anti-human CD3-PE-Cy5, 1:100; anti-human CD56-BUV737, 1:200; anti-human β2m-APC, 1:200; and anti-human HLA class I-FITC, 1:200.
For each sample, 50 μl solution is required. To account for volume loss during pipetting, it is recommended to prepare 10% to 20% excess solution.
2.Transfer 1 × 105 siRNA-treated NK cells to a 96-well conical microwell plate with lid.
3.Add PBS to each well to a final volume of 200 μl.
4.Centrifuge 5 min at 300 × g , room temperature.
5.Remove supernatant.
6.Add 50 μl staining solutions to the appropriate cell pellets.
7.Mix well and incubate for 20 min at room temperature in the dark.
8.Wash cells twice in PBS:
-
For the first wash, add 150 μl PBS to the wells. Centrifuge 5 min at 300 ×g, room temperature, and remove supernatant.
-
For the second wash, resuspend cell pellet in 200 μl PBS. Centrifuge 5 min at 300 ×g, room temperature, and remove supernatant.
9.Resuspend cell pellet in 100 μl fixation buffer.
10.Incubate for 15 min at 4°C in the dark.
11.Wash cells twice in FACS buffer:
-
For the first wash, add 150 μl FACS buffer to the wells. Centrifuge 5 min at 500 ×g, room temperature, and remove supernatant.
-
For the second wash, resuspend cell pellet in 200 μl FACS buffer. Centrifuge 5 min at 500 ×g, room temperature, and remove supernatant.
12.Resuspend cell pellet in 100 μl FACS buffer.
13.Analyze sample by flow cytometry.
14.Use analysis software to evaluate the expression of β2m and HLA class I protein on viable CD3– CD56+ NK cells.
REAGENTS AND SOLUTIONS
Cryostorage medium
- 90% (v/v) fetal bovine serum (FBS; e.g., Sigma-Aldrich, cat. no. F7524-500ML)
- 10% (v/v) DMSO (e.g., Sigma-Aldrich, cat. no. D2650-100ML)
- Sterilize through a 0.22-μm filter
- Store at –20°C for up to 4 weeks
Culture medium
- RPMI-1640 (e.g., Sigma-Aldrich, cat. no. R0883-500ML) containing:
- 10% (v/v) FBS (e.g., Sigma-Aldrich, cat. no. F7524-500ML)
- 2 mM L-glutamine (e.g., Sigma-Aldrich, cat. no. 59202C-100ML)
- 100 U/ml penicillin (e.g., Gibco, cat. no. 15140122)
- 100 ug/ml streptomycin (e.g., Gibco, cat. no. 15140122)
- Sterilize through a 0.22-μm filter
- Store at 4°C for up to 2 weeks
FACS buffer
- PBS, pH 7.4 (e.g., Gibco, cat. no. 10010023) containing:
- 2 mM EDTA (e.g., Invitrogen, cat. no. AM9260G)
- 2% (v/v) FBS (e.g., Sigma-Aldrich, cat. no. F7524-500ML)
- Sterilize through a 0.22-μm filter
- Store at 4°C for up to 4 weeks
MACS buffer
- PBS, pH 7.4 (e.g., Gibco, cat. no. 10010023) containing:
- 2 mM EDTA (e.g., Invitrogen, cat. no. AM9260G)
- 0.5% (v/v) FBS (e.g., Sigma-Aldrich, cat. no. F7524-500ML)
- Sterilize through a 0.22-μm filter
- Store at 4°C for up to 4 weeks
COMMENTARY
Background Information
Several protocols for genetically manipulating NK cells with various methodologies have been developed for the study of gene functions. Generally, NK cells are considered challenging to transfect, and workflows using electroporation, lipofection, or viral transduction result in variable delivery efficiency or can cause impaired viability (reviewed in Carlsten & Childs, 2015). Protocols using CRISPR/Cas9 have been described with varying efficiency. Lambert, Leijonhufvud, Segerberg, Melenhorst, & Carlsten (2020) achieved editing efficiencies of 50% when targeting CXCR4 in in vitro expanded NK cells. Rautela, Surgenor, & Huntington (2020) obtained 24% deletion of NKp46 in resting NK cells and up to 90% deletion following in vitro activation, and similar efficiencies have been reported for ADAM17 , PDCD1 , and CISH (Pomeroy et al., 2020). In contrast to these available approaches, the experimental procedure detailed here does not require in vitro activation or expansion of NK cells for efficient delivery of siRNA. Chemically modified siRNA enables passive uptake without electroporation or other techniques that reduce viability and has been demonstrated to achieve near complete uptake in NK cells while maintaining viability (Palacios et al., 2022).
This article describes the treatment of primary human NK cells with commercially available siRNA molecules and details all steps required from isolation of human PBMCs from buffy coats to evaluation of knockdown efficiency.
The described technique has been effectively applied to disrupt protein expression of several genes of interest including B2M , TGFBR2 , CXCR1 , and CX3CR1 (Palacios et al., 2022).
Critical Parameters
Before enrichment of NK cells, it is critical to maintain high viability in PBMC samples. When working with cryopreserved PBMCs, it is strongly advised to follow the described resting period to allow for optimal recovery of viable cells.
Following enrichment of NK cells, any excess protein from MACS buffer (or culture medium if the cells were cultured after enrichment) should be removed. Presence of excess protein can impede the passive uptake procedure and impair delivery of siRNA, thereby reducing the degree of RNA interference in the treated cells.
The siRNA treatment procedure is a simple and short workflow, but the selection of the employed siRNA reagents and the appropriate techniques to evaluate knockdown are of outmost importance. As such, characterizing the gene of interest and its expression dynamics in primary human NK cells before manipulation studies represents a critical parameter. Furthermore, it is recommended to validate obtained findings with complementary techniques to exclude off-target effects (Lin et al., 2019).
Troubleshooting
High viability of NK cells during siRNA delivery is crucial for successful experimentation. We have provided detailed instructions for procedures to obtain NK cells with high viability from human PBMCs and have emphasized a resting period before NK cell enrichment when working with cryopreserved PBMCs. In case the viability of the enriched NK cells is high directly before siRNA treatment but drops after starting the treatment, it is recommended to titrate the concentration of recombinant human IL-15 that is supplemented in the siRNA delivery medium. Specific subsets of NK cells or specific applications might require higher doses of IL-15, and the biological activity of the cytokine may vary. One option to provide enhanced survival signals could be to add recombinant cytokine again after a certain culture period (e.g., add cytokine again after 24 hr of treatment). Alternatively, other prosurvival factors for NK cells such as recombinant human IL-2 can be tested. To exclude that changes in viability are on-target effects that are triggered by knockdown of the gene of interest, it is recommended to assess viability in conditions treated with nontargeting control siRNA.
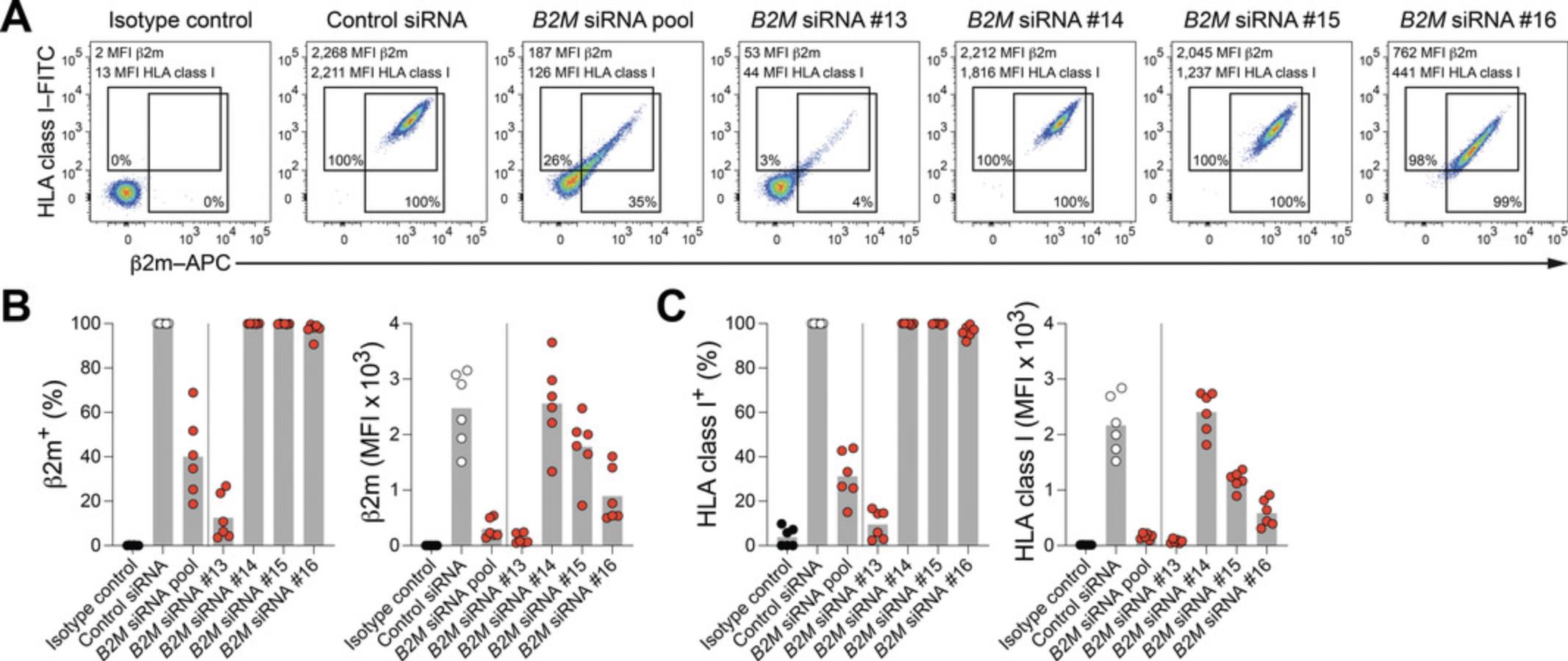
Low knockdown efficiency is an anticipated potential pitfall and will strongly depend on the target gene. The protocol described herein has been demonstrated to achieve decent knockdown effects across multiple genes of interest (Palacios et al., 2022). However, targeting novel genes may require additional optimization of the process. To exclude that delivery of siRNA molecules is the bottleneck that causes low knockdown efficiency, it is highly recommended to assess the uptake efficiency with a fluorescently labeled nontargeting siRNA (e.g., FAM-labeled Accell Green Nontargeting Control siRNA; Horizon Discovery, cat. no. D-001950-01-20). Furthermore, it is recommended to longitudinally evaluate knockdown efficiency of every novel target gene at both the transcript and the protein level (e.g., by reverse-transcription quantitative PCR and flow cytometry, respectively). Strongly affected expression at the transcript but not protein level might be due to high protein stability. In such a scenario, prolonged treatment time or repeated treatment may be required to achieve reduced protein expression. Comparing changes at the transcript and protein level over treatment time will also facilitate determining optimal timing for analysis. Moreover, as illustrated in Figure 2, different individual siRNA sequences that target distinct regions of the gene of interest may result in significantly different knockdown efficiencies. It is thus recommended to either use SMARTpool products that contain four individual sequences or test all available individual sequences in parallel and select the best performing reagent.
Anticipated Results
Figure 1 displays the purity of viable CD14– CD19– CD3– CD56+ NK cells after enrichment from human PBMCs (Basic Protocol 1).
Figure 2 provides an example of targeting the B2M gene in NK cells with different siRNA reagents (SMARTpool mix of four siRNA sequences or individual siRNA sequences 13, 14, 15, and 16) after 96 hr of treatment (Basic Protocol 3). Flow cytometric evaluation of β2m protein expression demonstrates efficient knockdown upon treatment with B2M siRNA SMARTpool and especially with B2M siRNA 13. These results illustrate the expected knockdown efficiency and also highlight a considerable degree of variability between individual siRNA sequences that target distinct regions of the B2M transcript. It is expected that silencing results vary depending on the gene of interest.
Time Considerations
As described in the protocol, enrichment of NK cells from human PBMCs (Basic Protocol 1) requires ∼1 hr. Preparation of siRNA (Basic Protocol 2) takes about 45 min, and the workflow to deliver siRNA into enriched NK cells (Basic Protocol 3) requires about 30 min to complete. The siRNA treatment period can span a period of 48 to 96 hr, depending on knockdown kinetics of the targeted gene and the assessment at transcript or protein level.
Acknowledgments
This project has received funding from the European Union's Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No. 838909 (to Q.H.) and H2020-MSC-ITN-765104-MATURE-NK (to K.-J.M.) This work was further supported by Knut and Alice Wallenberg Foundation (KAW 2018-011 to K.-J.M.), Swedish Research Council (2020-02286 to K.-J.M.), Swedish Children's Cancer Society (PR2020-0159 to K.-J.M.), Swedish Cancer Society (CAN 2017/676 to K.-J.M.), Research Council of Norway (275469 to K.-J.M.), Norwegian Cancer Society (223310 to K.-J.M.), National Institutes of Health (NIH; PTE Federal Award No: P01CA111412, Subaward No: P009500901 to K.-J.M.), South-Eastern Norway Regional Health Authority (2021073 to K.-J.M.), Region Stockholm (2020-0733 to P.M.), Petrus och Augusta Hedlunds Stiftelse (M2021-1533 and M2022-1821 to Q.H.), Stiftelsen Clas Groschinskys Minnesfold (M21120 and M2233 to Q.H.), Stiftelsen Torspiran (to Q.H.), and Stiftelsen Lars Hiertas Minne (FO2021-0263 to Q.H.).
Author Contributions
Pouria Momayyezi : formal analysis, investigation, writing—review and editing; Karl-Johan Malmberg : supervision, writing—review and editing, funding acquisition; Quirin Hammer : conceptualization, formal analysis, investigation, supervision, visualization, writing—original draft, writing—review and editing, funding acquisition.
Conflict of Interest
K.-J.M. is a scientific advisor for Fate Therapeutics and Vycellix and has received research funding from Fate Therapeutics. The remaining authors declare no conflicts of interest.
Open Research
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Literature Cited
- Bjorkstrom, N. K., Strunz, B., & Ljunggren, H. G. (2022). Natural killer cells in antiviral immunity. Nature Reviews Immunology , 22(2), 112–123. doi: 10.1038/s41577-021-00558-3
- Carlsten, M., & Childs, R. W. (2015). Genetic manipulation of NK cells for cancer immunotherapy: Techniques and clinical implications. Frontiers in Immunology , 6, 266. doi: 10.3389/fimmu.2015.00266
- Chiossone, L., & Vivier, E. (2022). Bringing natural killer cells to the clinic. Journal of Experimental Medicine , 219, e20220830. doi: 10.1084/jem.20220830
- Lambert, M., Leijonhufvud, C., Segerberg, F., Melenhorst, J. J., & Carlsten, M. (2020). CRISPR/Cas9-based gene engineering of human natural killer cells: Protocols for knockout and readouts to evaluate their efficacy. Methods in Molecular Biology , 2121, 213–239. doi: 10.1007/978-1-0716-0338-3_18
- Lin, A., Giuliano, C. J., Palladino, A., John, K. M., Abramowicz, C., Yuan, M. L., … Sheltzer, J. M. (2019). Off-target toxicity is a common mechanism of action of cancer drugs undergoing clinical trials. Science Translational Medicine , 11, eaaw8412. doi: 10.1126/scitranslmed.aaw8412
- Palacios, D., Momayyezi, P., Huhn, O., Ask, E. H., Dunst, J., Malmberg, K. J., & Hammer, Q. (2022). An optimized platform for efficient siRNA delivery into human NK cells. European Journal of Immunology , 52, 1190–1193. doi: 10.1002/eji.202149710
- Pomeroy, E. J., Hunzeker, J. T., Kluesner, M. G., Lahr, W. S., Smeester, B. A., Crosby, M. R., … Moriarity, B. S. (2020). A genetically engineered primary human natural killer cell platform for cancer immunotherapy. Molecular Therapy , 28, 52–63. doi: 10.1016/j.ymthe.2019.10.009
- Rautela, J., Surgenor, E., & Huntington, N. D. (2020). Drug target validation in primary human natural killer cells using CRISPR RNP. Journal of Leukocyte Biology , 108, 1397–1408. doi: 10.1002/JLB.2MA0620-074R
- Wolf, N. K., Kissiov, D. U., & Raulet, D. H. (2022). Roles of natural killer cells in immunity to cancer, and applications to immunotherapy. Nature Reviews Immunology , [Epub ahead of print]. doi: 10.1038/s41577-022-00732-1
Key Reference
- Palacios et al. (2022). See above.
- Reports a simple and efficient method to deliver siRNA into primary human NK cells and demonstrates its versatility with various target genes.

