Cassava Metabolomics and Starch Quality
Alisdair R. Fernie, Paul D. Fraser, Paul D. Fraser, Laise Rosado-Souza, Laise Rosado-Souza, Laure C. David, Laure C. David, Margit Drapal, Margit Drapal, Jörg Hofmann, Jörg Hofmann, Patrick A. W. Klemens, Patrick A. W. Klemens, Frank Ludewig, Frank Ludewig, H. Ekkehard Neuhaus, H. Ekkehard Neuhaus, Toshihiro Obata, Toshihiro Obata, Laura Perez-Fons, Laura Perez-Fons, Armin Schlereth, Uwe Sonnewald, Mark Stitt, Samuel C. Zeeman, Wolfgang Zierer
Abstract
Cassava plays an important role as a staple food for more than 800 million people in the world due to its ability to maintain relatively high productivity even in nutrient-depleted soils. Even though cassava has been the focus of several breeding programs and has become a strong focus of research in the last few years, relatively little is currently known about its metabolism and metabolic composition in different tissues. In this article, the absolute content of sugars, organic acids, amino acids, phosphorylated intermediates, minerals, starch, carotenoids, chlorophylls, tocopherols, and total protein as well as starch quality is described based on multiple analytical techniques, with protocols specifically adjusted for material from different cassava tissues. Moreover, quantification of secondary metabolites relative to internal standards is presented using both non-targeted and targeted metabolomics approaches. The protocols have also been adjusted to apply to freeze-dried material in order to allow processing of field harvest samples that typically will require long-distance transport. © 2019 The Authors.
Basic Protocol 1 : Metabolic profiling by gas chromatography–mass spectrometry (GC-MS)
Support Protocol 1 : Preparation of freeze-dried cassava material
Support Protocol 2 : Preparation of standard compound mixtures for absolute quantification of metabolites by GC-MS
Support Protocol 3 : Preparation of retention-time standard mixture
Basic Protocol 2 : Determination of organic acids and phosphorylated intermediates by ion chromatography–mass spectrometry (IC-MS)
Support Protocol 4 : Preparation of standards and recovery experimental procedure
Basic Protocol 3 : Determination of soluble sugars, starch, and free amino acids
Alternate Protocol : Determination of soluble sugars and starch
Basic Protocol 4 : Determination of anions
Basic Protocol 5 : Determination of elements
Basic Protocol 6 : Determination of total protein
Basic Protocol 7 : Determination of non-targeted and targeted secondary metabolites
Basic Protocol 8 : Determination of carotenoids, chlorophylls, and tocopherol
Basic Protocol 9 : Determination of starch quality
INTRODUCTION
Metabolomics is a constant evolving technology for biochemical analysis of complex mixtures, with the current aim of allowing unbiased, quantitative analysis of very large numbers of metabolites, if not all, in a biological-material extract (Hall & Hardy, 2012). The plant metabolome is not only highly complex but also highly dynamic. The physical and chemical properties of metabolites render them highly diverse, and therefore, no single extraction process and no single analytical method is able to access all metabolites (Fernie, Trethewey, Krotzky, & Willmitzer, 2004). In general, metabolites in plants can be separated into primary and secondary/specialized metabolites (Fernie & Pichersky, 2015). Primary metabolites, such as sugars, amino acids, and organic acids, serve as intermediates required for anabolic and catabolic energy metabolism and are found in most free-living plants. By contrast, the so-called secondary/specialized metabolites comprise several thousands of individual compounds that are usually much less abundant than primary metabolites and mainly serve ecological functions. Additionally, several metabolic pathways are influenced by inorganic cations, and a shortage or excess in different tissues may directly or indirectly limit plant growth and development.
Cassava is an important staple food for more than 800 million people in Africa, Latin America, and Asia (Chetty, Rossin, Gruissem, Vanderschuren, & Rey, 2013; De Souza et al., 2017; Howeler, Lutaladio, & Thomas, 2013) due to its favorable adaptation to diverse soils and environmental conditions and its compatibility with a variety of farming systems with minimum agricultural inputs (Chandrasekara & Josheph Kumar, 2016). Cassava's roots are an important source of carbohydrates (mainly starch), whereas its leaves are high in protein and additionally provide a supply of vitamins and minerals (Montagnac, Davis, & Tanumihardjo, 2009). Cassava is also characterized by high content of toxic cyanogenic glucosides, and especially linamarin, with significant amounts accumulating in all organs, including the leaves, stems, and roots (Picmanova et al., 2015). The role of cyanogenic glycosides as defense compounds relies on their bioactivation by specific endogenous β-glycosidases, causing release of hydrogen cyanide (HCN), which is toxic to respiring organisms (Bjarnholt et al., 2018). For this reason, linamarin must be removed before dietary use by humans or as feed. Although cassava became a focus of research in the last few years, especially in regard to disease susceptibility and post-harvest deterioration, we still lack a comprehensive understanding of the metabolic characteristics of and the huge variability. In this article, we describe most of the more standard and best-developed analytical approaches to dissect the chemical composition of cassava material at a tissue level. Basic Protocols 1 and 3 allow absolute quantification of sugars, organic acids, amino acids, and additional metabolites specific for each protocol, whereas Basic Protocol 2 focuses on organic acids and phosphorylated metabolites. The four support protocols describe preparation of freeze-dried material (Support Protocol 1) and standard compounds (Support Protocols 2 to 4). Basic Protocols 4, 5, and 6 respectively center on anions, elements, and total protein content essential for determining the nutritional level of a plant. In addition, Basic Protocol 7 permits relative quantification of secondary metabolites as a result of non-targeted and targeted metabolomics approaches. Determination of carotenoids, chlorophylls, and tocopherol is described in Basic Protocol 8. In addition to the starch quantification described in Basic Protocol 3 and the Alternate Protocol, Basic Protocol 9 presents detailed analysis of starch quality. The combination of such techniques lays the groundwork for further studies by providing robust procedures that allow a very diverse overview of the complexity of the metabolism and metabolic composition of cassava.
Basic Protocol 1: METABOLIC PROFILING BY GAS CHROMATOGRAPHY–MASS SPECTROMETRY (GC-MS)
Metabolic profiling using GC-MS allows high accuracy and sensitivity in the analysis of highly complex mixtures of compounds, such as plant tissue samples, permitting identification and robust quantification of metabolites from a single extract. GC-MS has relatively extensive coverage of compounds involved in the central pathways of primary metabolism, such as sugars, sugar alcohols, amino acids, organic acids, and polyamines. It is the most widely used technique for plant metabolomics, and solid protocols for machine set-up, sample preparation, analysis, and chromatogram evaluation have been developed.
The protocol presented here is carried out essentially as previously described by Lisec, Schauer, Kopka, Willmitzer, & Fernie (2006), with modifications specific for freeze-dried cassava tissue samples (Support Protocol 1). Metabolite profiles are in general expressed as ratios, with data compared with a control sample. This protocol can also be used for absolute quantification when authentic standards with different amounts of each compound are included in each batch of samples.
Materials
-
Homogenized, ground, and freeze-dried cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
100% methanol (gradient grade for liquid chromatography; Merck, CAS 67-56-1)
-
Ribitol (Sigma, CAS 488-81-3)
-
Chloroform (CHCl3; for liquid chromatography; Merck, CAS 67-66-3)
-
Ultrapure (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Argon 5.0 (Messer-Griesheim) or another inert gas
-
Standard compound mixtures (for absolute quantification of metabolites by GC-MS; see Support Protocol 2)
-
20 mg/ml methoxyamine hydrochloride (Sigma, CAS 593-56-6) in pyridine (Merck, CAS 110-86-1)
-
MSTFA (N -methyl-N -(trimethylsilyl)trifluoroacetamide, Macherey and Nagel, CAS 24589-78-4)
-
Fatty-acid methyl esters (FAMEs; see Support Protocol 3)
-
Helium 5.0 carrier gas
-
Analytical scale
-
2.0-ml screw-cap tubes (e.g., Sarstedt, 72.693)
-
70°C Thermomixer (e.g., Eppendorf Thermomixer comfort) or thermostable water bath
-
Microcentrifuge
-
Glass tubes with screw caps (e.g., 6-ml tubes, DURAN, 231751159)
-
Vortex
-
Standard tabletop centrifuge
-
1.5-ml tubes (e.g., Eppendorf Safe-Lock microcentrifuge tubes)
-
Long pipet tips (200-µl volume; MBP KLP pipet tips, 3540-11)
-
Vacuum concentrator (e.g., SpeedVac, Thermo Fischer Scientific)
-
Air-flow sample concentrator (optional)
-
37°C Thermomixer (e.g., Eppendorf Thermomixer comfort)
-
1.1-ml GC-MS sample vials with screw-top lids (e.g., Chromatographie Zubehör Trott, 45 11 00 740)
-
Gas chromatograph (Agilent, 6890N)
-
Split and splitless injector with electronic pressure control up to 150 psi (Agilent)
-
Autosampler system (PAL, Agilent)
-
Pegasus III GC-time-of-flight/MS (GC-TOF/MS) mass analyzer (LECO)
-
Capillary column (MDN-35, 30 m × 0.32 mm, 0.25-mm film thickness; Supelco or equivalent)
-
Conical single-taper split/splitless liner (Agilent)
-
Chroma TOF 1.0 software (LECO) or equivalent
Metabolic profiling extraction
1.Weigh out 10 mg (±1 mg) of each homogenized, ground, and freeze-dried cassava tissue in a 2.0-ml screw-cap tube. Prepare an additional empty tube as a blank.
2.Make a mixture of 1400 µl of 100% methanol and 60 µl ribitol (stock concentration of 0.2 mg/ml) per sample. Pre-cool to −20°C.
3.Pipet 1460 µl cold methanol-ribitol solution onto each cassava tissue sample.
4.Shake at 1000 rpm for 15 min at 70°C using a Thermomixer or thermostable water bath.
5.Centrifuge 10 min at 20,000 × g.
6.Transfer supernatant to a glass tube with a screw cap.
7.Add 750 µl chloroform.
8.Add 1500 µl ultrapure water and thoroughly vortex for 15 s.
9.Centrifuge 15 min at 4000 × g.
10.Using long pipet tips, transfer two 200-µl aliquots from upper phase (polar phase) into 1.5-ml tubes.
11.Transfer six 200-µl aliquots from blank tube into fresh 1.5-ml tubes.
12.Dry aliquots of samples as well as blanks and standard compound mixtures in a vacuum concentrator for ≥3 hr (or overnight), without heating. Immediately after drying the aliquots, flush tubes with argon 5.0 or another inert gas and close tubes tightly. Store tubes at −80°C until derivatization.
Derivatization (ONLY the polar phase)
13.Place dry aliquots of samples as well as blanks and standard compound mixtures stored at −80°C in a vacuum concentrator or air-flow sample concentrator for 30 min before derivatization.
14.Add 40 µl of 20 mg/ml methoxyamine hydrochloride in pyridine to the dry aliquots (samples, blanks, and quantitative controls).
15.Shake at 950 rpm for 2 hr at 37°C using a Thermomixer.
16.Spin down briefly to collect drops on the cap into the main bulk.
17.Prepare MSTFA mix (1 ml MSTFA + 20 µl FAMEs) and add 70 µl to each tube.
18.Shake at 950 rpm for 30 min at 37°C using a Thermomixer.
19.Spin down briefly to collect drops on the cap.
20.Transfer contents into a 1.1-ml GC-MS sample vial with a screw-top lid.
GC-TOF/MS
21.Using a gas chromatograph, a split and splitless injector with electronic pressure control up to 150 psi, an autosampler system, and a Pegasus III GC-TOF/MS mass analyzer, inject 1 µl sample into a capillary column at 230°C in splitless and split (1:30 ratio) modes using a conical single-taper split/splitless liner, with the helium 5.0 carrier gas flow set to 2 ml/min. Keep flow rate constant with electronic pressure control enabled. Include syringe-washing steps before and after each injection in injection program.
22.Perform chromatography using a temperature program of isothermal for 2 min at 80°C, followed by a 15°C/min ramp to 330°C, holding at this temperature for 6 min, and then cooling as rapidly as instrument specifications allow. Set transfer-line and ion-source temperatures to 250°C.
23.Set recorded mass to be m/z 70 to 600 scanning range at 20 scans/s. Set remaining monitored chromatography time to proceed with a 180-s solvent delay, with filaments turned off. Set manual mass defect to 0, filament bias current to −70 V, and detector voltage to about 1700 to 1850 V. Tune instrument automatically according to the manufacturer's instructions.
24.Use Chroma TOF 1.0 software or equivalent for visualization of chromatograms and export of .cdf files compatible with further analysis. Create a list of average retention times of each FAME, using mass 87 as a reference, for calculation of the retention index (RI) and correction of each chromatogram separately in subsequent steps.
25.Perform target analysis of cassava metabolites.
26.Once the data matrix has been exported for the samples and standards, correct values relative to the internal standard, ribitol, by dividing each metabolite value by the ribitol value in each sample. To generate standard curves, plot values for the standard mixes and the amounts of metabolites used. Calculate absolute values for each metabolite from the curves, with final values divided by the sample weight.
Support Protocol 1: PREPARATION OF FREEZE-DRIED CASSAVA MATERIAL
Freeze-drying (lyophilization or cryodesiccation) is a dehydration process at low temperature and pressure. It is an excellent method for preserving material after the material has been harvested and snap-frozen in liquid nitrogen. Several protocols in this article (Basic Protocols 1 to 9 and Alternate Protocol) have been adjusted to apply to freeze-dried material in order to allow processing of field harvest samples that typically require long-distance transport. Freeze-dried material can be transported at room temperature and weighs significantly less than non-freeze-dried material, making it more convenient.
Materials
-
Cassava tissue (e.g., source leaves, sink leaves, upper stems, and/or roots)
-
Liquid nitrogen
-
Dry ice (optional)
-
Cutting tool (optional)
-
Polyvials (e.g., liquid nitrogen–safe 20-ml polyvials, Zinsser Analytic, 3071401) or equivalent container for harvest
-
Freeze drier with vacuum pump (e.g., Christ, model Alpha 2-4)
-
Analytical scale
-
Mixer mill (Retsch, model MM 300, https://www.retsch.com)
-
Steel balls (e.g., 5- to 7-mm diameter, Th. Geyer, https://www.thgeyer.com/en/)
-
Paper tissue (e.g., Kimwipes, KIMTECH Science, 107281)
-
Elastic bands
-
Vacuum bags
-
Desiccator
1.Harvest appropriate cassava tissue according to the experimental aim.
2.For thick materials such as storage roots, cut tissue into small pieces before placing it into a polyvial or equivalent container for harvest.
3.Immediately freeze tissue in liquid nitrogen and store at −80°C until further processing.
4.Turn on and set up freeze drier with a vacuum pump in advance according to the manufacturer's instructions.
5.Place sample on dry ice or in a container of liquid nitrogen. Prepare sample for the freeze drier by removing the lid and then quickly place it into freeze drier.
6.Freeze-dry until completely dry. To verify if the sample is fully dried, weigh sample periodically until a constant weight is noted.
7.Grind sample using a mixer mill and steel balls until a fine powder forms. During the grinding process, keep sample at low temperature by periodically cooling with liquid nitrogen or dry ice.
8.Freeze-dry ground sample again (see steps 4 to 6) for ≥2 days to ensure complete removal of water before aliquoting. To avoid loss or cross-contamination of the material during this process, remove lid and cover container with paper tissue fixed with an elastic band. Close container with the corresponding lid immediately after removing it from the freeze drier.
9.Aliquot material according to amount specified in the chosen protocol (Basic Protocols 1 to 9 or Alternate Protocol).
10.For long-term storage, store material under vacuum in vacuum bags in a desiccator at 4°C.
Support Protocol 2: PREPARATION OF STANDARD COMPOUND MIXTURES FOR ABSOLUTE QUANTIFICATION OF METABOLITES BY GC-MS
Metabolite profiles are in general expressed as ratios, with data compared with a control sample. However, absolute quantification is possible when authentic standards are included, allowing generation of standard curves (Basic Protocol 1, step 26). Here, we present the complete list of metabolites that we have used for absolute quantification of cassava metabolites. These metabolites are divided into two mixes to avoid overlap of specific metabolites due to similar retention times.
Materials
-
Standard metabolite solutions (1 mg/ml each; see Table 1)
-
2.5:1 methanol/water solution (methanol gradient grade for liquid chromatography; Merck, CAS 67-56-1; ultrapure water, ∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
0.2 mg/ml ribitol (Sigma, CAS 488-81-3)
-
Argon 5.0 (Messer-Griesheim) or another inert gas
-
15-ml Falcon tubes
-
1.5-ml microcentrifuge tubes
-
Vacuum concentrator (e.g., SpeedVac, Thermo Fischer Scientific)
| Mix | Metabolite | GMDa ref. no. | ChEBIb ID | Molecular weight (g/mol) | CAS no. | Supplier | Cat. no. |
|---|---|---|---|---|---|---|---|
| 1 | 2-Oxoglutaric acid | R002922 | 30915 | 146.11 | 328-50-7 | Sigma | K1128 |
| 1 | Alanine | R000072 | 16449 | 89.1 | 56-41-7 | Sigma | A7627 |
| 1 | Arginine | R003205 | 29016 | 174.2 | 7200-25-1 | Sigma | A8094 |
| 1 | Asparagine | R000314 | 22653 | 132.12 | 70-47-3 | Sigma | A0884 |
| 1 | Aspartate | R000384 | 22660 | 133.1 | 56-84-8 | Sigma | A8949 |
| 1 | β-Alanine | R001368 | 16958 | 89.09 | 107-95-9 | Sigma | A7752 |
| 1 | Citrate | R000112 | 30769 | 192.12 | 77-92-9 | Sigma | C4540 |
| 1 | Erythritol | R000642 | 17113 | 122.12 | 149-32-6 | Supelco | R-422470 |
| 1 | Fructose | R001682 | 28757 | 180.2 | 57-48-7 | Sigma | F0127 |
| 1 | Fumarate | R000659 | 18012 | 116.1 | 110-17-8 | Sigma | F2752 |
| 1 | 4-Amino-butyric acid (GABA) | R003031 | 16865 | 103.12 | 56-12-2 | Sigma | A5835 |
| 1 | Galactinol | R003201 | 17505 | 342.3 | 16908-86-4 | Sigma | 79544 |
| 1 | Glucose | R001393 | 17234 | 180.2 | 50-99-7 | Sigma | G8270 |
| 1 | Glutamate | R000687 | 18237 | 147.1 | 56-86-0 | Sigma | G1251 |
| 1 | Glutamine | R000689 | 28300 | 146.15 | 56-85-9 | Sigma | G3126 |
| 1 | Glycerol | R000680 | 17754 | 92.09 | 56-81-5 | Sigma | G5516 |
| 1 | Glycerol-3P | R002017 | 15978 | 172.014 | 57-03-4 | Wako | 198-02052 |
| 1 | Glycine | R000681 | 15428 | 75.1 | 56-40-6 | Sigma | G7126 |
| 1 | Histidine | R001410 | 27570 | 155.16 | 4998-57-6 | Sigma | 53330 |
| 1 | Isoleucine | R000732 | 24898 | 131.2 | 73-32-5 | Sigma | I2752 |
| 1 | Leucine | R003029 | 25017 | 131.2 | 61-90-5 | Sigma | L8000 |
| 1 | Lysine | R000063 | 25094 | 146.2 | 923-27-3 | Sigma | L8021 |
| 1 | Malate | R000108 | 6650 | 134.1 | 97-67-6 | Sigma | M6413 |
| 1 | Methionine | R003032 | 16811 | 149.21 | 63-68-3 | Sigma | M9625 |
| 1 | Myo-inositol | R000105 | 17268 | 180.2 | 87-89-8 | Sigma | I5125 |
| 1 | Ornithine | R000464 | 18257 | 132.16 | 3184-13-2 | Sigma | O4386 |
| 1 | Phenylalanine | R002940 | 28044 | 165.2 | 673-06-3 | Sigma | P1751 |
| 1 | Phosphate | 26078 | 97.99 | 7664-38-2 | Carl Roth | 6366 | |
| 1 | Proline | R001417 | 26271 | 115.13 | 344-25-2 | Sigma | 81705 |
| 1 | Putrescine | R000182 | 17148 | 88.2 | 333-93-7 | Sigma | P7505 |
| 1 | Pyruvate | R003170 | 32816 | 88.1 | 113-24-6 | Sigma | P8574 |
| 1 | Raffinose | R003156 | 16634 | 504.42 | 17629-30-0 | Sigma | R0250 |
| 1 | Serine | R001418 | 17822 | 105.1 | 312-84-5 | Sigma | 84970 |
| 1 | Succinate | R000114 | 15741 | 118.1 | 110-15-6 | Sigma | S3674 |
| 1 | Sucrose | R000523 | 17992 | 342.3 | 57-50-1 | Sigma | S9378 |
| 1 | Trehalose | R002802 | 16551 | 342.3 | 6138-23-4 | Sigma | T9531 |
| 1 | Tryptophan | R000539 | 27897 | 204.22 | 73-22-3 | Sigma | T0254 |
| 1 | Tyrosine | R000511 | 18186 | 181.19 | 60-18-4 | Sigma | T3754 |
| 1 | Valine | R000541 | 27266 | 117.2 | 72-18-4 | Sigma | V0500 |
| 1 | Xylose | R000094 | 18222 | 150.13 | 58-86-6 | Sigma | X1500 |
| 2 | Benzoate | R000565 | 30746 | 122.12 | 65-85-0 | Sigma | 242381 |
| 2 |
3-Trans-caffeoylquinic acid (chlorogenic acid) |
R000125 | 16112 | 354.31 | 327-97-9 | Sigma | C3878 |
| 2 | Cis-caffeic acid | 17395 | 180.16 | 331-39-5 | Sigma | C0625 | |
| 2 | Fucose | R000096 | 33984 | 164.16 | 3615-37-0 | Sigma | F8150 |
| 2 | Galactonolactone | R000122 | 15895 | 178.14 | 1668-08-2 | Sigma | 5313 |
| 2 | Glycerate | R000686 | 33508 | 106.1 | 6057-35-8 | Sigma | 50032 |
| 2 | Glycolate | R000683 | 17497 | 76.05 | 79-14-1 | Sigma | G8284 |
| 2 | Homoserine | R000037 | 30653 | 119.12 | 1927-25-9 | Sigma | H6515 |
| 2 | Lactate | R000802 | 28358 | 90.1 | 10326-41-7 | Sigma | L0625 |
| 2 | Linamarin | R000410 | 16441 | 247.248 | 9008-22-4 | Sigma | L9634 |
| 2 | Maleate | R000433 | 18300 | 116.07 | 110-16-7 | Sigma | 92816 |
| 2 | Maltose | R000438 | 17306 | 342.3 | 6363-53-7 | Sigma | M5885 |
| 2 | N-acetyl-serine | R000039 | 45441 | 147.129 | 97-14-3 | Sigma | A2638 |
| 2 | Nicotinate | R000542 | 15940 | 123.1 | 59-67-6 | Merck | 481918 |
| 2 | O-acetyl-serine | R000042 | 17981 | 147.129 | 66638-22-0 | Sigma | A6262 |
| 2 | 4-Hydroxy-cinnamate (p-coumaric acid) | R000198 | 32374 | 164.047 | 501-98-4 | Sigma | 55823 |
| 2 | Quinic acid | R000474 | 26493 | 192.17 | 77-95-2 | Sigma | 138622 |
| 2 | Rhamnose | R000104 | 26546 | 164.16 | 6155-35-7 | Sigma | R3875 |
| 2 | Shikimate | R003041 | 16119 | 174.15 | 138-59-0 | Sigma | S5375 |
| 2 | Threonate | R002093 | 15908 | 136.1 | 7306-96-9 | Sigma | 380644 |
| 2 | Threonine | R000538 | 26986 | 119.12 | 80-68-2 | Sigma | T8625 |
| 2 | Trans-caffeic acid | R001373 | 16433 | 180.16 | 331-39-5 | Sigma | 60018 |
| 2 | Urea | R002926 | 16199 | 60.06 | 57-13-6 | Sigma | U5378 |
- a
GMD, Golm Metabolome Database (http://gmd.mpimp-golm.mpg.de/).
- b
ChEBI, Chemical Entities of Biological Interest (https://www.ebi.ac.uk/chebi/).
Mix 1 (40 compounds)
1.Add 250 µl of each standard metabolite solution for Mix 1 to a 15-ml Falcon tube (total volume of 10,250 µl).
2.Bring volume to 12,500 µl with 2.5:1 methanol/water solution (final concentration of 20 ng/µl for each compound in mix).
3.Make a 1:10 dilution (2 ng/µl) of mix.
4.Make aliquots of volumes indicated in Table 2 into 1.5-ml microcentrifuge tubes according to final concentration.
| Final amount (ng) | Volume (µl) | Dilution (stock concentration) |
|---|---|---|
| 250 | 125 | 1:10 (2 ng/µl) |
| 500 | 250 | 1:10 (2 ng/µl) |
| 1000 | 50 | Not diluted (20 ng/µl) |
5.Add 4 µl of 0.2 mg/ml ribitol to each tube (final amount of 0.8 µg).
6.Dry contents of the tubes in a vacuum concentrator overnight. Immediately after drying the aliquots, flush tubes with argon 5.0 or another inert gas and close tubes tightly. Store at −80°C until derivatization.
Mix 2 (23 compounds)
7.Add 250 µl of each standard metabolite solution for Mix 2 to a 15-ml Falcon tube (total volume of 10,250 µl).
8.Bring volume to 10,000 µl with 2.5:1 methanol/water solution (final concentration of 20 ng/µl for each compound in mix).
9.Make a 1:10 dilution (1:10,200 ng/µl) of mix.
10.Make aliquots of volumes indicated in Table 3 into 1.5-ml microcentrifuge tubes according to final concentration.
| Final amount (ng) | Volume (µl) | Dilution (stock concentration) |
|---|---|---|
| 250 | 100 | 1:10 (2 ng/µl) |
| 500 | 200 | 1:10 (2 ng/µl) |
| 1000 | 40 | Not diluted (20 ng/µl) |
11.Repeat steps 5 and 6 for Mix 2.
Support Protocol 3: PREPARATION OF RETENTION-TIME STANDARD MIXTURE
Basic Protocol 1 uses FAMEs as retention-time standards for calculation of the RI. The RI is a crucial parameter for peak assignment and for avoiding false peak annotations (Lisec et al., 2006). Due to expected variance in absolute retention times, each chromatogram must be corrected for retention times separately using the RI. Alternatively to FAMEs, alkanes or fatty acids can be used. All must be of standard grade for GC.
Materials
-
FAMES (see Table 4)
-
Chloroform (CHCl3; for liquid chromatography; Merck, CAS 67-66-3).
-
1.5-ml microcentrifuge tubes
-
Vortex
-
50-ml Falcon tube
-
Glass vials
| Name | Formula | C-chain | CAS no. | Sigma cat. no. | Phase | Density | Mass (mg) or volume (µl) |
|---|---|---|---|---|---|---|---|
| Methylcaprylate | C9H18O2 | C8 | 111-11-5 | 21719-1ML-F | liquid | 0.887 | 20 |
| Methylpelargonate | C10H20O2 | C9 | 1731-84-6 | 76368-1ML | liquid | 0.884 | 20 |
| Methylcaprate | C11H22O2 | C10 | 110-42-9 | 21479-1ML | liquid | 0.881 | 20 |
| Methyllaurate | C13H26O2 | C12 | 111-82-0 | 234591-2.5G | liquid | 0.88 | 20 |
| Methylmyristate | C15H30O2 | C14 | 124-10-7 | 70129-1ML | liquid/warm up | 0.866 | 20 |
| Methylpalmitate | C17H34O2 | C16 | 112-39-0 | 76159-1G | solid | 20 | |
| Methylstearate | C19H38O2 | C18 | 112-61-8 | 85769-1G | solid | 10 | |
| Methyleicosanoate | C21H42O2 | C20 | 1120-28-1 | 10941-1G | solid | 10 | |
| Methyldocosanoate | C23H46O2 | C22 | 929-77-1 | 11940-1G | solid | 10 | |
| Lignoceric acid methylester | C25H50O2 | C24 | 2442-49-1 | 87115-250MG | solid | 10 | |
| Methylhexacosanoate | C27H54O2 | C26 | 5802-82-4 | 52203-100MG | solid | 10 | |
| Methyloctacosanoate | C29H58O2 | C28 | 55682-92-3 | 74701-250MG | solid | 10 | |
| Triacontanoic acid methylester | C31H62O2 | C30 | 629-83-4 | 63642-100MG | solid | 10 |
1.Add 10 or 20 mg or 10 or 20 µl of each FAME into individual 1.5-ml microcentrifuge tubes.
2.Bring volume to 1 ml with chloroform and mix by vortexing.
3.Combine all solutions (liquids and dissolved solids) into a 50-ml Falcon tube and bring up to a final volume of 25 ml with chloroform.
4.Aliquot into glass vials and store at −20°C until use as specified in Basic Protocol 1, step 17.
Basic Protocol 2: DETERMINATION OF ORGANIC ACIDS AND PHOSPHORYLATED INTERMEDIATES BY ION CHROMATOGRAPHY–MASS SPECTROMETRY (IC-MS)
IC-MS combined with perchloric-acid extraction is a powerful method for targeted or untargeted analysis of ionic compounds such as highly unstable phosphorylated metabolites and organic acids, e.g., of glycolysis, the Calvin cycle, or the Krebs cycle, from plant and other tissues. The method avoids derivatization and, in combination with a triple quadrupole MS, does not need good chromatographic separation for most metabolites. The extraction protocol presented is optimized for cassava leaf and root tissue and is derived from our previously described method (Hofmann, Bornke, Schmiedl, Kleine, & Sonnewald, 2011).
Materials
-
Freeze-dried cassava tissue or frozen and powdered cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
Liquid nitrogen
-
1 M perchloric acid [70% (v/v) perchloric acid (Sigma-Aldrich, CAS 7601-90-3) diluted with distilled water at 1:11.7], 4°C
-
0.1 M perchloric acid (1 M perchloric acid diluted with distilled water at 1:10), 4°C
-
5 M potassium carbonate (K2CO3; Sigma-Aldrich, CAS 584-08-7), 4°C
-
Distilled water
-
100 mM potassium hydroxide (KOH)
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Analytical scale
-
2-ml safe-lock reaction tubes (Sarstedt, 72.691)
-
Mortars and pestles
-
Timer
-
4°C microcentrifuge
-
pH indicator paper strips (Carl Roth, 0549.2)
-
10-kDa multiwell filter plates (AcroPrep Advance, Omega 10K MWCO, Pall, 8034) or 10-kDa spin columns (Pall, OD010C34)
-
300-µl conical plastic vials (VWR, 548-0120) with snap ring caps/Teflon lids (VWR, 548-0016)
-
High-performance liquid chromatography (HPLC) system (ICS-3000, Thermo Scientific Dionex), including eluent generator, autosampler, KOH suppressor system, and conductivity detector driven by Chromeleon 6.8 software and DCMSLink software
-
HPLC columns: IonPac® AG11-HC guard column (2 × 50 mm, Dionex) and two AS11-HC columns (2 × 250 mm, Thermo Scientific Dionex) in series
-
Triple quadrupole mass spectrometer (QTrap 3200, Applied Biosystems) with Analyst 1.6.2 software
Sample extraction
1.Weigh out 25 mg (±3 mg) freeze-dried cassava tissue or 100 mg (±3 mg) frozen and powdered cassava tissue per 2-ml safe-lock reaction tube. Store tubes in liquid nitrogen or at −80°C until use.
2.Label three 2-ml safe-lock reaction tubes for each sample/extract and pre-cool on ice.
3.Place four samples in a container of liquid nitrogen.
4.Cool two mortars with inserted pestles by filling with ∼50 ml liquid nitrogen each.
5.Start timer.
6.When the liquid nitrogen has evaporated from the mortars, add one sample and 1 ml cold 1 M perchloric acid to each mortar. Thoroughly homogenize each sample using the pestle until the sample begins to thaw.
7.After ∼7 min, cool two additional mortars and pestles with liquid nitrogen (see step 4).
8.When the liquid nitrogen has evaporated, add one sample and 1 ml cold 1 M perchloric acid to each mortar. Thoroughly homogenize each sample using the pestle until the sample begins to thaw.
9.When the sample is homogenous and thawed, pipet suspension into a pre-cooled 2-ml reaction tube on ice (see step 2) and rinse residual homogenate from mortar and pestle with 1 ml cold 0.1 M perchloric acid. Pool with initial 1-ml suspension, resulting in 2 ml plant extract per sample.
10.Centrifuge four extracts for 2 min at 18,700 × g , 4°C.
11.Transfer 1.8 ml supernatant into a new pre-cooled 2-ml reaction tube on ice (see step 2) and immediately add 50 µl of 5 M K2CO3.
12.Add another 50 µl of 5 M K2CO3 and wait a few seconds, until CO2 evaporation/bubbling is low. Close tubes.
13.Invert four extracts three times, let them sit for ∼15 s, and then open lids with caution due to pressure buildup. Check that pH is between 6.8 and 7.2 with a pH indicator paper strip. If the pH is still acidic, add small volumes (5 to 20 µl) of 5 M K2CO3 and recheck pH.
14.Centrifuge four extracts for 2 min at 18,700 × g , 4°C, to remove precipitated potassium chlorate.
15.Transfer supernatant to a new pre-cooled 2-ml reaction tube on ice (see step 2) and store at −80°C until analysis.
16.Repeat steps 4 to 15 for next four samples. Repeat as needed until all samples have been processed.
17.Prior to IC-MS analysis, filter extracts through 10-kDa multiwell membrane filter plates or 10-kDa spin columns: Wet filters with 5 µl distilled water. Add 200 µl extract to each well and centrifuge 15 min at 15,300 × g , 4°C.
18.Add 80 µl filtered, bubble-free extract to a 300-µl conical plastic vial with a snap ring cap/Teflon lid.
IC-MS
19.Using an HPLC system and columns, set the following chromatography parameters:
-
Distilled water (eluent A) with changes of 100 mM KOH (eluent B) within a total time of 80 min; flow rate, 0.25 ml/min; column temperature, 35°C.
-
Injection volume: 10 µl.
-
Autosampler temperature: 4°C.
-
Gradient for eluent B: 0 min, 4%; 0-1 min, 4%; 1-6 min, 15%; 6-12 min, 19%; 12-22 min, 20%; 22-24 min, 23%; 24-27 min, 35%; 27-37 min, 38%; 37-39 min, 45%; 39-44 min, 100%; 44-71 min, 100%; 71-76 min, 4%; and 76-80 min, 4%.
20.Set suppressor current gradient to follow the KOH gradient starting at 4 mA, with a maximum of 100 mA.
21.To minimize contamination, set flow to the mass spectrometer to between 5 and 52 min only. Conduct a blank run (10 µl ultrapure-water injection) of 40 min in between samples with the following gradient for eluent B: 0 min, 4%; 0.1-10 min, 35%; 10-15 min, 100%; 15-44 min, 100%; 44-50 min, 4%; and 50-55 min, 4%.
22.Perform QTrap detection with a triple quadrupole mass spectrometer in multiple reaction monitoring (MRM) mode with the following settings:
-
Scan ranges: fromm/z87-606 (precursor ions) andm/z59-385 (product ions)
-
ESI-source parameters: −4500 V, 600°C.
-
N2-gas pressures: 20 psi (curtain gas), 30 psi (gas 1), 20 psi (gas 2), and collision gas set to medium.
-
Dwell time for ions: 75 ms.
-
Compound-specific potentials for recorded ions (see Table5) and ion spectra determined by infusion and tuning with standard solutions of 1-10 mM.
-
Scan time per cycle: 3.7 s.
| Metabolite | Q1a (mass) | Q3a (mass) | Dwell time (msec) | DPb | EPb | CEPb | CEb | CXPb |
|---|---|---|---|---|---|---|---|---|
| Pyruvate | 87 | 59 | 75 | −20 | −4.5 | −14 | −12 | −4 |
| G1P | 259 | 79 | 75 | −30 | −4 | −30 | −20 | 0 |
| G6P | 259 | 97 | 75 | −25 | −5 | −30 | −20 | 0 |
| F6P | 259 | 97 | 75 | −25 | −5 | −30 | −20 | 0 |
| M6P | 259 | 97 | 75 | −25 | −5 | −30 | −20 | 0 |
| 3PG | 185 | 97 | 75 | −15 | −8 | −16 | −18 | −4 |
| UDPglc | 565 | 323 | 75 | −50 | −6 | −40 | −30 | −4 |
| ADPglc | 588 | 346 | 75 | −45 | −7 | −33.412 | −30 | −4 |
| G16BP | 339 | 97 | 75 | −20 | −4 | −24.199 | −26 | −4 |
| F16BP | 339 | 159 | 75 | −30 | −4.5 | −24.199 | −40 | −4 |
| PEP | 167 | 79 | 75 | −15 | −4 | −17.835 | −18 | −4 |
| Cit | 191 | 87 | 75 | −15 | −2.5 | −18.723 | −26 | −4 |
| Icit | 191 | 111 | 75 | −25 | −4 | −18.723 | −16 | −4 |
| Mal | 133 | 115 | 75 | −20 | −4 | −16.577 | −12 | −4 |
| AMP | 346 | 79 | 75 | −35 | −7 | −24.458 | −50 | −4 |
| ADP | 427 | 79 | 75 | −25 | −7 | −27.455 | −72 | −4 |
| ATP | 427 | 159 | 75 | −25 | −7 | −27.455 | −36 | −4 |
| Ru15P2 | 310 | 97 | 75 | −20 | −4 | −20 | −24 | 0 |
| UDP | 404 | 79 | 75 | −35 | −7 | −26.604 | −90 | −4 |
| S6P | 421 | 97 | 75 | −45 | −8 | −27.233 | −40 | −2 |
| T6P | 421 | 241 | 75 | −45 | −8 | −27.233 | −40 | −2 |
| DHAP | 169 | 97 | 75 | −15 | −2 | −18 | −14 | 0 |
| aKG | 145 | 101 | 75 | −15 | −3 | −10 | −14 | 0 |
| Succ | 117 | 73 | 75 | −20 | −6 | −18 | −16 | 0 |
| E4P | 199 | 97 | 75 | −20 | −7.5 | −19.019 | −16 | 0 |
| Fum | 115 | 71 | 75 | −15 | −6.5 | −15.911 | −10 | 0 |
| UDPNAG | 606.3 | 385 | 75 | −65 | −8 | −34.089 | −36 | −4 |
| Shik | 173 | 111 | 75 | −25 | −5.5 | −18.057 | −14 | 0 |
- a
Q1, quadrupole; Q3, fragment ions.
- b
Potentials are in volts: declustering, DP; entrance, EP; collision cell entrance, CEP; collision, CE; and collision cell exit, CXP.
23.Calculate metabolite content of samples with Chromeleon software from recorded peak areas for precursor/product-ion transitions and comparison to known standard curves.
Support Protocol 4: PREPARATION OF STANDARDS AND RECOVERY EXPERIMENTAL PROCEDURE
In this protocol, purchasable analytical-grade metabolites are used for standard-curve generation and validation recovery experiments for Basic Protocol 2.The individual metabolites are quantified via their respective areas under the HPLC peak curve. For precise quantification, different dilutions of the standard mixture should be included in measurements at the beginning, at the end, and approximately every 10 to 20 samples. Spike-in experiments are performed in order to check for extraction losses and matrix effects. Because not every metabolite is extracted with equal efficiency and because metabolites may behave differently during measurement, every metabolite should be checked for its extraction and measurement efficiency by adding a known amount of the analytically pure substance prior to extraction. By calculating the ratio of measured substance to input substance, the efficiency can be determined for each metabolite, and accuracy of measurement can be ensured.
Additional Materials (also see Basic Protocol 2)
- Analytical-grade metabolites (∼5 mg each; e.g., from Sigma-Aldrich)
Standard mix preparation
1.Prepare 100 mM stock solutions of analytical-grade metabolites in distilled water for the following low-abundance metabolites: adenosine triphosphate (ATP), adenosine diphosphate (ADP), adenosine monophosphate (AMP), uridine diphosphate (UDP), adenosine diphosphate–glucose (ADP-Glc), UDP-glucose (UDP-Glc), UDP-N- acetylglucosamine (UDP-GlcNAc), pyruvate, phosphoenolpyruvate (PEP), erythrose-4-phosphate (E4P), dihydroxyacetone phosphate (DHAP), pyrophosphate (PPi), ribulose-1,5-bisphosphate (Ru15BP), glucose-1-phosphate (G1P), glucose-1,6-bisphosphate (G16BP), fructose-1,6-bisphosphate (F16BP), sucrose-6-phosphate (S6P), trehalose-6-phosphate (T6P), trehalose, and mannose-6-phosphate (M6P).
2.Prepare 500 mM stock solutions of analytical-grade metabolites in distilled water for the following high-abundance metabolites: citrate (Cit), isocitrate (ICit), malate (Mal), glucose-6-phosphate (G6P), fructose-6-phosphate (F6P), 3-phosphoglyceric acid (3PG), fumarate (Fum), alpha-ketoglutarate (aKG), succinate, and shikimate.
3.Mix equal volumes of each stock solution to make a “standard mix” containing all 30 metabolites, with a concentration of either 3.33 mM (low-abundance metabolites) or 16.66 mM (high-abundance metabolites).
| Name | Dilution (in dH2O) | Final concentrationa |
|---|---|---|
| Standard mix | None | 16.66 mM/3.33 mM |
| STD1 | 1:100 of standard mix | 166.60 µM/33.30 µM |
| STD2 | 1:300 of standard mix | 55.53 µM/11.10 µM |
| STD3 | 1:10 of STD1 | 16.66 µM/3.33 µM |
| STD4 | 1:10 of STD2 | 5.55 µM/1.11 µM |
| STD5 | 1:10 of STD3 | 1.66 nM/0.33 nM |
| STD6 | 1:10 of STD4 | 0.55 nM/0.11 nM |
| STD7 | 1:10 of STD5 | 0.16 nM/0.03 nM |
- a
High-abundance metabolites/low-abundance metabolites.
Validation recovery experiments
4.Add a defined amount of metabolite of interest (approximately 1 to 2 times the measured content in the tissue; use a purchasable analytical-grade metabolite standard) to the frozen-tissue powder (freeze-dried cassava tissue or frozen and powdered cassava tissue) prior to extraction.
5.Extract metabolites from the spiked tissue and from unspiked control tissue and measure metabolite concentrations in the samples according to Basic Protocol 2.
6.Calculate theoretical value for the metabolite of interest in the spiked sample by adding the defined amount of the spiked metabolite of interest (see step 4) to the value measured in the unspiked sample. Divide measured value for the metabolite of interest in the spiked sample by the theoretical value to determine the recovery rate.
Basic Protocol 3: DETERMINATION OF SOLUBLE SUGARS, STARCH, AND FREE AMINO ACIDS
Generally, sugars are water soluble and fuel various metabolic pathways with carbon and energy. Thus, they can be converted to other carbohydrates, like starch or cellulose, or they provide substrates for main metabolic routes, like glycolysis or the oxidative pentose phosphate pathway. These two pathways generate precursors for amino-acid synthesis, making plants able to be fully photoautotrophic. The levels of sugars, starch, and amino acids are thus, on the one hand, indicative of the metabolic state of cells or tissues and, on the other hand, influence plant growth, organ development, and overall yield. Therefore, it is of great importance to gain knowledge on the levels of sugars, starch, and amino acids in various cassava tissues under selected conditions.
Here, we describe an easy-to-follow protocol that can be used for a large number of samples in a short period of time and that allows extraction of soluble sugars and amino acids in one extraction. Extraction and determination of aspartate, glutamate, serine, glutamine, threonine, alanine, glycine, cysteine, proline, valine, lysine, phenylalanine, arginine, histidine, tyrosine, methionine, isoleucine, and leucine via HPLC have been tested for various tissues, although cysteine and methionine, depending on the type of sample and genotype, are sometimes not detectable. The extraction and digestion of starch were optimized for high-starch-containing tissues types, e.g., cassava storage roots.
Materials
-
Homogenized and fine-milled freeze-dried cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
80% (v/v) ethanol (p.a., absolute; CHEMSOLUTE, 2246.2500, CAS 64-175)
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Starch digestion mix (see recipe; make fresh)
-
Borate buffer: 0.2 M boric acid (H3BO3; p.a.; Merck, 1.00165.1000), pH 8.8 (adjust pH with sodium hydroxide, or NaOH)
-
AQC stock solution: 2 mg/ml AQC (6-aminoquinolyl-N- hydroxysuccinimidyl carbamate; C14H11N3O4; Synchem, CAS 148757-94-2) in acetonitrile (HPLC gradient grade; Fisher Scientific, CAS 75-05-8)
-
Eluent A: 100 mM sodium acetate and 7 mM triethanolamine in ultrapure water, pH 5.2
-
Eluent B: 90% (v/v) acetonitrile and 10% (v/v) ultrapure water
-
Amino Acid Standard H (Thermo Fisher Scientific, 20088)
-
2.5 mM glutamine
-
Analytical scale
-
1.5-ml reaction tubes with screw caps
-
Vortex (e.g., Vortex-Genie 2)
-
37°C and 80°C Thermomixers (e.g., Thermomixer comfort, Eppendorf)
-
4°C or room-temperature microcentrifuge (e.g., Eppendorf Centrifuge 5417 R)
-
Autoclave
-
55°C and 95°C heating blocks
-
Glass vials for HPLC (TECHLAB, FLK 103.040) with lids (TECHLAB, FLK 101.227), septa (TECHLAB, FLK 102.797), and micro-inserts (TECHLAB, FLK 101.263)
-
HPLC system: Dionex P680-HPLC system with ASI-100 Automated Sample Injector (Dionex), RF 2000 fluorescence detector (Dionex), column system consisting of CC8/4 ND 100-5 C18ec and CC 250/4 ND 100-5 C18ec (Macherey-Nagel), and Chromeleon 6.7 software
-
Additional reagents and equipment for glucose assay (see Alternate Protocol, steps 11 to 21)
Sample preparation
1.Weigh out 5 mg (±0.5 mg) homogenized and fine-milled freeze-dried cassava tissue in a 1.5-ml reaction tube with a screw cap.
2.Add 0.5 ml of 80% ethanol and vortex thoroughly.
3.Shake at 500 rpm for 30 min at 80°C using a Thermomixer.
4.Store sample for 10 min on ice.
5.Centrifuge 10 min at 16,000 × g , 4°C.
6.Carefully transfer supernatant into a new 1.5-ml reaction tube with a screw cap. Perform glucose assay according to Alternate Protocol, steps 11 to 21.
7.Dissolve pellet in 0.5 ml of 80% ethanol.
8.Centrifuge 10 min at 16,000 × g , 4°C.
9.Carefully remove supernatant.
10.Wash pellet in 1 ml ultrapure water.
11.Centrifuge 10 min at 16,000 × g , 4°C.
12.Remove and discard supernatant.
13.Dissolve pellet in 0.5 ml ultrapure water.
14.Autoclave at 121°C for 20 min.
15.Cool down sample to room temperature and proceed with starch digestion.
Starch digestion
16.Prepare starch digestion mix for the number of samples plus one.
17.Add 0.5 ml starch digestion mix to autoclaved pellet (see step 15).
18.Vortex briefly.
19.Shake at 200 rpm (mild agitation) for ≥4 hr at 37°C.
20.Inactivate enzymes at 95°C for 5 min using a heating block.
21.Cool down sample to room temperature.
22.Centrifuge 10 min at 16,000 × g , room temperature.
23.Transfer supernatant into a new 1.5-ml reaction tube and discard pellet. Perform glucose assay according to Alternate Protocol, steps 11 to 21.
Derivatization and HPLC analysis of amino acids
24.Mix 20 µl ethanol extraction (see step 6) with 60 µl borate buffer and 20 µl AQC stock solution in a 1.5-ml reaction tube.
25.Vortex immediately for 10 s.
26.Incubate at 55°C for 10 min using a heating block.
27.Transfer solution into a glass vial for HPLC with a lid, septum, and micro-insert.
28.Determine amino acids via HPLC system with the following chromatography parameters:
-
Eluent A: 100 mM sodium acetate and 7 mM triethanolamine in ultrapure water, pH 5.2.
-
Eluent B: 90% acetonitrile and 10% ultrapure water.
-
Flow rate, 1 ml/min; column temperature, 32°C.
-
Injection volume: 20 µl.
-
Gradient for eluent B: 0 min, 0%; 2 min, 3%; 28 min, 8%; 28.5 min, 11%; 31 min, 12%; 35 min, 17%; 42 min, 24%; 47 min, 30%; 47 min, 100%; 52 min, 100%; 52 min, 0%; and 61 min, 0%.
-
Detection of AQC amino acids by fluorescence with excitation at 254 nm and emission at 395 nm.
Preparation and HPLC analysis of amino-acid standards
29.Mix 10 µl Amino Acid Standard H with 10 µl of 2.5 mM glutamine and 30 µl ultrapure water in a 1.5-ml reaction tube.
30.Combine 10 µl of the mixture with 10 µl ultrapure water, 60 µl borate buffer, and 20 µl AQC stock solution.
31.Vortex immediately for 10 s.
32.Incubate at 55°C for 10 min using a heating block.
33.Transfer standard solution into a glass vial for HPLC with a lid, septum, and micro-insert.
34.For standard curves, measure 20, 10, and 5 µl standard solution via HPLC system.
35.For calculation of the total concentration of each amino acid, consider all dilutions made during procedure (i.e., AQC derivatization at 1:5, 20-µl injection volume from 500-µl extraction volume).
36.To convert the raw concentration values obtained from the HPLC system (in nmol/20 µl injection volume) into mg/g DW, perform the following calculation:
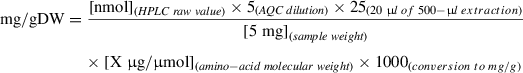
Alternate Protocol: DETERMINATION OF SOLUBLE SUGARS AND STARCH
In order to quantify starch and sugars, perchloric-acid extraction is ideal, as all enzymes that may metabolize carbohydrates are immediately inactivated. Additionally, compared to ethanol extraction (Basic Protocol 3), fewer pigments get extracted alongside sugars, leading to lower background during further quantification. The extraction protocol presented is optimized for cassava tissues and is derived from our previously described method (Hostettler et al., 2011). Starch (in the digested pellet) and sucrose, glucose, and fructose (in the supernatant) can then be quantified with an enzyme-based glucose assay (Fig. 1).
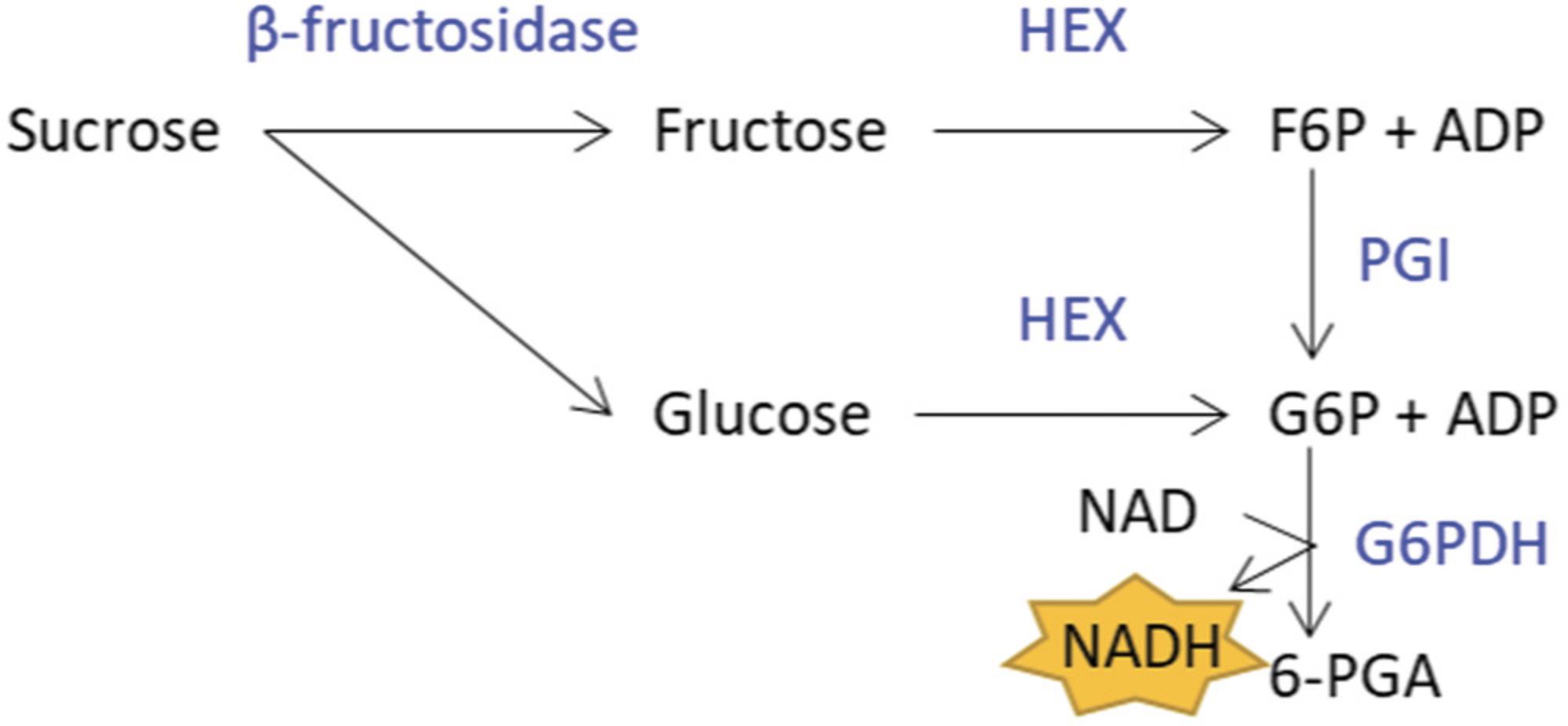
Materials
-
Freeze-dried cassava tissue or frozen and powdered cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
0.7 M perchloric acid [4.2% (v/v), prepared from 11.6 M/70% perchloric acid, Sigma, CAS 7601-90-3), 4°C
-
Deionized water
-
2 M KOH (Sigma, CAS 1310-58-3)/0.4 M MES (Sigma, CAS 145224-94-8), 4°C
-
80% (v/v) and 100% ethanol (puriss. p.a.), 4°C
-
Starch digestion mix (see recipe; make fresh)
-
Glucose assay mix (see recipe)
-
Glucose 6-phosphate dehydrogenase (G6PDH; grade II, from Leuconostoc mesenteroides , 1000 U/ml; Roche, 10165875001)
-
Phosphoglucomutase (PGI; from yeast, 2 mg/ml; Roche, 10127396001)
-
β-fructosidase (from yeast, 75,0000 U/2g; Roche, 104922)
-
Analytical scale
-
1.5- and 2-ml reaction tubes
-
Vortex
-
4°C microcentrifuge (e.g., Eppendorf Centrifuge 5417 R)
-
96-deep-well plate (1.3-ml volume)
-
95°C water bath
-
37°C incubator
-
Standard tabletop centrifuge
-
96-well flat-bottom microtiter plate
-
Microtiter plate reader
-
Multichannel pipet
Sample preparation
1.Weigh out 25 mg freeze-dried cassava tissue or 50 to 100 mg frozen and powdered cassava tissue per 2-ml reaction tube.
2.Add 0.5 ml ice-cold 0.7 M perchloric acid to tube and vortex thoroughly.
3.Incubate on ice for 30 min.
4.Centrifuge 10 min at 18,000 × g , 4°C.
5.Carefully transfer supernatant into a 1.5-ml reaction tube. Store on ice.
6.Add 0.5 ml deionized water to pellet and vortex to resuspend.
7.Centrifuge 10 min at 18,000 × g , 4°C.
8.Pool supernatant with that of first spin (see step 5). Leave tube containing the pellet on ice and proceed with supernatant.
9.Process supernatant:
- Add small aliquots (5 µl) of ice-cold 2 M KOH/0.4 M MES to supernatant until the pH reaches 7.0. Note volume added to neutralize.
Do this slowly and do not overshoot. Around 200 µl is usually needed.
- Centrifuge 10 min at 18,000 × g, 4°C.
Assay the supernatant immediately or freeze at −80°C and assay later.
10.Process pellet:
-
Wash pellet with 1 ml ice-cold 100% ethanol. Resuspend pellet by vortexing.
-
Centrifuge 5 min at 18,000 ×g, 4°C. Discard supernatant.
-
Wash pellet two additional times with 1 ml ice-cold 80% ethanol.
For green tissues, add one or two extra washes, until the supernatant is almost clear, to get rid of most of the chlorophyll.
-
After discarding the last wash, leave pellet to dry on a bench for 20 min.
-
Resuspend pellet in 1 ml deionized water.
-
Place two 50-µl aliquots of starch suspension per sample in a 96-deep-well plate.
Use wide-orifice pipet tips.
For starch quantification, one aliquot will be digested, and the other one will be a control.
Samples can be stored at −80°C at this point.
- Boil plate by floating it in a 95°C water bath for 20 min.
Make sure that the plate is properly sealed with the appropriate lid.
- Cool down plate to room temperature and spin briefly to remove condensed water and residual insoluble material from the lid.
Avoid cooling to below room temperature, in which case the starch will re-form secondary structures (retrogradation).
- Add 50 µl starch digestion mix to each well. Vortex briefly.
During this step, starch is hydrolyzed to glucose using amyloglucosidase and α-amylase. Also include an undigested control (starch suspension and 200 mM sodium acetate/acetic acid, pH 4.8) to assess any glucose contamination, which may occur if the pellet was not washed properly.
- Incubate at 37°C for 4 hr using an incubator.
It is critical that the starch is completely digested. Check the digestion efficiency by pipetting aliquots of the digest and control suspensions into microcentrifuge tubes containing Lugol solution (Carl Roth, N052.2). The digested sample should not stain for starch, whereas the control sample will.
- Centrifuge 10 min at 4000 ×g, room temperature.
Assay immediately for glucose or freeze at −20°C and assay later.
Glucose assay
11.Mix appropriate volume of sample with glucose assay mix to achieve a final volume of 200 µl per well in a 96-well flat-bottom microtiter plate.
12.Record absorbance at 340 nm using a microtiter plate reader.
13.Add 0.45 U G6PDH to each well using a multichannel pipet.
14.Record absorbance at 340 nm after a constant value is reached.
15.Add 0.3 µl PGI to each well using a multichannel pipet.
16.Record absorbance at 340 nm after a constant value is reached.
17.Dissolve ∼5 mg β-fructosidase in 100 µl deionized water and add 1 µl to each well using a multichannel pipet.
18.Record absorbance at 340 nm after a constant value is reached.
19.Plot absorbance of the standards against the amount of glucose. Determine linear range and calculate slope of the standard curve.
20.Calculate glucose content of the starch samples using the slope of the standard curve and the following ΔOD values and taking into account dilutions made during the procedure:
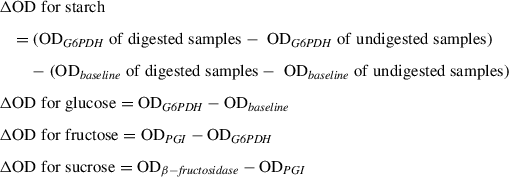

21.Express starch, fructose, and sucrose quantities in mol of C6 equivalent (glucose)/g of tissue or convert these values into mg/g of tissue by taking into account the molecular weights (glucose and fructose: 180 g/mol; sucrose: 342 g/mol; for starch, 1 µmol glucose is equivalent to 162 µg starch).
Basic Protocol 4: DETERMINATION OF ANIONS
IC is a powerful tool to determine the nutritional level of a plant. Depending on soil types, the availability of anions, especially like nitrate, phosphate, and sulfate, and their uptake into and distribution within the plant provide proxy information on the physiological status of corresponding tissues. The following protocol was tested for different freeze-dried and fresh samples derived from several cassava tissues.
Materials
-
Homogenized and fine-milled freeze-dried cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Anion Standard Mixed Concentrate (Fisher Chemical, J/4552)
-
1.8 mM sodium carbonate/1.7 mM sodium hydrogencarbonate
-
50 mM sulfuric acid
-
Analytical scale
-
1.5-ml reaction tubes with screw caps
-
95°C Thermomixer (e.g., Eppendorf Thermomixer comfort)
-
Microcentrifuge (e.g., Eppendorf Centrifuge 5417 R)
-
Vials for IC (TECHLAB, FLK 101.697) with lids/caps (TECHLAB, FLK 103.149)
-
IC system: 761-IC compact device (Metrohm), Dynamax automatic sample injector AI-1A (Rainin/Varian), column system consisting of Metrosep A Supp 4/5 Guard 4.0 and Metrosep A Supp 4-250/4.0 (Metrohm), and IC Net V2.3 Metrodata software (Metrohm)
1.Weigh out 5 mg (±0.5 mg) homogenized and fine-milled freeze-dried cassava tissue in a 1.5-ml reaction tube with a screw cap.
2.Pipet 0.5 ml ultrapure water into tube and screw on cap.
3.Shake at 600 rpm for 15 min at 95°C using a Thermomixer.
4.Remove sample from the Thermomixer and let it cool down to room temperature.
5.Centrifuge 10 min at 16,000 × g , room temperature.
6.Transfer supernatant into a new 1.5-ml reaction tube.
7.Transfer 100 µl sample or Anion Standard Mixed Concentrate into a vial for IC, close vial with a lid/cap, and put vial into the autosampler.
8.Perform chromatography using an IC system, separating anions using a Metrosep A Supp 4/5 Guard 4.0 and Metrosep A Supp 4-250/4.0 (Metrohm) column system, followed by conductometry of 20 µl sample at a flow rate of 1 ml/min and pressure of 7.5 MPa, using 1.8 mM sodium carbonate/1.7 mM sodium hydrogencarbonate as mobile phase and 50 mM sulfuric acid as counterion.
9.Prepare standard curves according to Table 7.
| Ion | STDa (1:10) | STD (5:6) | STD (2:3) | STD (1:2) | STD (1:3) |
|---|---|---|---|---|---|
| F- | 3 ppm | 2.5 ppm | 2 ppm | 1.5 ppm | 1 ppm |
| Cl- | 6 ppm | 5 ppm | 4 ppm | 3 ppm | 2 ppm |
| Br- | 25 ppm | 20.83 ppm | 16.6 ppm | 12.5 ppm | 8.3 ppm |
| NO3- | 30 ppm | 25 ppm | 20 ppm | 15 ppm | 10 ppm |
| PO43- | 60 ppm | 50 ppm | 40 ppm | 30 ppm | 20 ppm |
| SO42- | 30 ppm | 25 ppm | 20 ppm | 15 ppm | 10 ppm |
| To pipet | 150 µl anion standard + 1350 µl Milli-Q H2O | 250 µl STD (1:10) + 50 µl Milli-Q H2O | 200 µl STD (1:10) + 100 µl Milli-Q H2O | 150 µl STD (1:10) + 150 µl Milli-Q H2O | 100 µl STD (1:10) + 200 µl Milli-Q H2O |
- a
STD, standard.
10.To convert the raw concentration values obtained from the IC system (in mg/L) into mg/g DW, multiply value by the factor obtained from the sample weight/sample volume (e.g., 100) divided by 1000.
Basic Protocol 5: DETERMINATION OF ELEMENTS
Inorganic ions (e.g., K+, Mg2+, Zn2+, and Fe2+) are taken up by the roots and transported to leaves. Several metabolic pathways are strongly influenced by these ions, and a shortage of ions in leaves may limit growth and potentially block carbohydrate utilization. Thus, detailed ion profiling by inductively coupled plasma (ICP) analysis will contribute to a detailed assessment of source and sink interactions in cassava. Based on the protocol of Klaumann et al. (2011), the following protocol describes step-by-step determination of elements in cassava tissues.
Materials
-
Freeze-dried cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
Nitric acid (HNO3; 65% p.a.; Bernd Kraft, 05119.4100)
-
Hydrogen peroxide (H2O2; 30%; VWR Chemicals, 23622.298, CAS 7722-84-1)
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Single-element standards (CPI International):
- Calcium (Ca; S4400-10M91)
- Iron (Fe; S4400-1000261)
- Potassium (K; 4400-10M411)
- Magnesium (Mg; 4400-10M311)
- Manganese (Mn; S4400-1000321)
- Phosphorus (P; 4400-10M391)
- Sulfur (S; 4400-10M544)
- Zinc (Zn; S4400-1000681)
-
Analytical scale
-
PTFE microwave vessels with screw caps (PTFE-TFM Buckets for HPR-1000/10S, MLS GmbH)
-
Microwave (MLS ETHOS microwave system including HPR-1000/10S Rotor, MLS GmbH)
-
15-ml tubes (Greiner Bio-One, 188271)
-
ICP system [iCAP 6300 DUO ICP-OES system (Thermo Scientific) equipped with ASX-260 Autosampler (CETAC) and iTEVA iCAP 2.4.071 software]
1.Weigh out 20 to 30 mg freeze-dried cassava tissue in a PTFE microwave vessel with a screw cap.
2.Prepare a mixture of 5 ml HNO3, 2 ml H2O2, and 3 ml ultrapure water per sample (digestion mix).
3.Add 10 ml digestion mix to sample.
4.Screw on vessel cap, put vessel into the microwave, and start program (2 min, 800 W, 50°C; 15 min, 800 W, 150°C; 8 min, 1000 W, 210°C; 15 min, 700 W, 210°C; Stop) for digestion.
5.After the digestion, unscrew vessel cap and transfer solution into a 15-ml tube.
6.Rinse vessel with 2 ml ultrapure water and transfer solution into the same 15-ml tube.
7.Mix by inverting tube 8 times.
8.Determine elemental composition by performing ICP analysis using an ICP system with the parameters described in Table 8.See Table 9 for elements, wavelengths, and plasma views.
| Parameter | Setting |
|---|---|
| Pump tubing | Sample orange white |
| Drain white/white | |
| Spray chamber | Glass cyclonic |
| Nebulizer | Glass concentric |
| Center tube | 2.0 mm |
| Pump speed | 50 rpm |
| Nebulizer gas flow | 0.45 L/min |
| RF power | 1150 W |
| Element | Wavelength (nm) | View |
|---|---|---|
| Ca | 317.933 | Radial |
| Fe | 238.204 | Axial |
| K | 766.49 | Radial |
| Mg | 279.079 | Axial |
| Mn | 257.61 | Axial |
| P | 177.434 | Axial |
| S | 182.034 | Axial |
| Zn | 202.548 | Axial |
9.For preparation of the calibration standards, use high-purity single-element standards at final concentrations given in Table 10.
| Element | Concentration (mg/kg) | Volume for 100 ml (ml) |
|---|---|---|
| Ca | 600 | 6 |
| Fe | 1 | 0.02 |
| K | 400 | 4 |
| Mg | 50 | 0.5 |
| Mn | 1 | 0.02 |
| P | 50 | 0.5 |
| S | 100 | 1 |
| Zn | 1 | 0.02 |
| Add 30 ml HNO3 (65%) and 57.94 ml Milli-Q H2O |
10.For calculation of the total element amount in mg/g DW, multiply raw value obtained from the ICP system by the dilution factor (here, for a final volume of 12 ml, the factor is 12) and the factor of the sample weight (e.g., for 25 mg = 1 g/0.025 g = 40) divided by 1000.
Basic Protocol 6: DETERMINATION OF TOTAL PROTEIN
Determination of the protein content of different cassava tissues is often described/performed according to the Kjeldahl method. When using this method, based on protein digestion, subsequent determination of the resulting amino acids is time consuming and labor intensive. Here, we describe an extraction method with a “simple” buffer, which, combined with determination according to the Bradford assay, gives robust and reproducible results for a large number of samples. This method can be done in almost every laboratory and does not rely on expensive equipment, such as an HPLC system.
Materials
-
Homogenized and fine-milled freeze-dried cassava tissue (source leaves, sink leaves, upper stems, and/or roots; see Support Protocol 1)
-
Protein extraction buffer (see recipe)
-
Roti®-Quant (Carl Roth, K015.1), including 5× staining solution
-
Bovine serum albumin (BSA) standards (Pierce™ Bovine Serum Albumin Standard Ampules, 2 mg/ml, Thermo Scientific, 23209)
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
Analytical scale
-
1.5-ml reaction tubes with screw caps
-
Vortex (e.g., Vortex-Genie 2)
-
4°C Thermomixer (e.g., Eppendorf Thermomixer comfort)
-
4°C microcentrifuge (e.g., Eppendorf Centrifuge 5417 R)
-
Photometer (e.g., Eppendorf BioPhotometer)
1.Weigh out 5 to 10 mg (±0.5 mg) homogenized and fine-milled freeze-dried fine-milled cassava tissue in a 1.5-ml reaction tube with a screw cap.
2a. For cassava leaves : Add 1 ml protein extraction buffer.
2b. For cassava storage roots : Add 0.25 ml protein extraction buffer.
3.Vortex for 10 s.
4.Shake at 1200 rpm for 10 min at 4°C using a Thermomixer.
5.Vortex for 10 s.
6.Centrifuge 5 min at 1500 × g , 4°C.
7.Remove supernatant completely and transfer it into a new 1.5-ml reaction tube. Store on ice.
8.Repeat steps 2 to 7 at least one more time with pellet until no more protein is extracted.
9.Measure total protein in the supernatant according to the Bradford assay (Bradford, 1976) and add values of each extraction together.
10.For quantification of proteins and preparation of standards, follow manual for Roti®-Quant, with BSA standards in protein extraction buffer as the protein for the standard curve.
11.Based on the number of samples, prepare a master mix of 1× staining solution, with 200 µl of 5× staining solution (from Roti®-Quant) and 775 µl ultrapure water per sample.
12.Label fresh 1.5-ml reaction tubes according to samples.
13a. For cassava leaves : Pipet 975 µl of 1× staining solution into a labeled tube for each sample. Add 25 µl extracted protein and mix by vortexing. To prepare a blank, add 25 µl protein extraction buffer and mix by vortexing.
13b. For cassava storage roots : Pipet 950 µl of 1× staining solution into a labeled tube for each sample. Add 50 µl extracted protein and mix by vortexing. To prepare a blank, add 50 µl protein extraction buffer and mix by vortexing.
14.Using a photometer, measure OD595 of the standard solutions and samples against the blank after 15 min.
15.Plot OD595 of the standard solutions and the amount of protein used. Read amount of protein in the samples on the calibration curve (in µg/ml). Multiply this value by protein extraction buffer volume (in ml; e.g., 0.25 or 1 ml) divided by the sample weight (in mg).
Basic Protocol 7: DETERMINATION OF NON-TARGETED AND TARGETED SECONDARY METABOLITES
A universal method for optimal extraction of all plant secondary metabolites is presently not feasible. The complexity, physical and chemical diversity, dynamic range, instability, and turnover preclude a single extraction and analytical platform for secondary metabolites. The extraction method provided below enables fast and reliable detection of a broad range of chemical classes. Furthermore, the method facilitates analysis of the same extracts on multiple/complementary analytical platforms without the necessity of repeating the extraction process. The extraction is based on liquid-liquid partition of immiscible solvents of different polarities, which facilitates extraction of polar and semi-polar compounds separately from non-polar metabolites in a single extraction.
Chromatographic methods based on reverse-phase mode provide good resolution of cassava's most-cited secondary metabolites, such as cyanogenic glycosides, flavonoids, hydroxycinnamic acids, lignans, hydroxybenzoates, and glycosylated apocarotenoids (Drapal, Barros de Carvalho, Ovalle Rivera, Becerra Lopez-Lavalle, & Fraser, 2019). These compounds are amenable to detection with MS using electrospray ionization in either negative or positive mode. Numerous methods describing analysis of secondary metabolites can be found in the literature, but generally, chromatographic separation can be achieved using a linear gradient of acidified water and acetonitrile or methanol on a C18 analytical column.
Regarding quantification, a general approach is to use relative quantification of molecular features by normalizing against the internal standard. Typically, accurate quantification methods are applied after statistical assessment of metabolomics datasets, which generates a list of candidate molecular features significantly associated with the trait of interest. Once compounds are identified, accurate quantification can be performed using internal- or external-standard calibration curves.
Materials
-
Freeze-dried cassava tissue (leaves, stems, and/or roots; see Support Protocol 1)
-
LC-MS-grade water (Fisher Scientific, 10728098)
-
Methanol (HPLC grade; Fisher Scientific, M/4056/17)
-
Chloroform (CHCl3; Fisher Scientific, C/4920/17)
-
Nitrogen (N2) stream (optional)
-
0.2 mg/ml genistein in ethanol (Extrasynthese, 1372S)
-
Acidified water: LC-MS-grade water containing 0.01% to 0.1% (v/v) formic acid (LC-MS grade; Merck, 5.33002.0050)
-
Acetonitrile (LC-MS grade; VWR Chemicals, 83640.290)
-
TissueRuptor (Qiagen, TH-03) or blade grinder (Genie, Hinari, MB280) or other alternative to TissueRuptor
-
Screw-capped 2-ml polypropylene tubes
-
TissueLyser LT (Qiagen, 23.1001/05735)
-
Stainless-steel beads (5 mm; Qiagen, 69989)
-
1.5- and 2-ml microcentrifuge tubes
-
Vortex
-
Tube rotator equipped with sample holder (Stuart, SB3)
-
Aluminum foil
-
Microcentrifuge (Eppendorf, 5424)
-
2-mm syringe filters with 0.45-µm nylon membranes
-
1-ml syringes
-
Vacuum centrifuge (Genevac EZ-2 Plus, BPS Crowthorne; optional)
-
Conical glass inserts (250 µl, with polypropylene spring-type feet) mounted in screw-capped 2-ml glass vials (clear or amber glass)
-
Caps with PTFE/red silicone septa
-
LC-MS system with electrospray source and automated sampler
-
Reverse-phase C18 analytical column (100 or 50 × 2.1 mm, i.d. 1.8 µm)
Sample preparation/grinding
1a. For freeze-dried cassava leaves : Grind freeze-dried tissue using a TissueRuptor.
1b. For more sturdy tissue, such as cassava stems and roots : Grind freeze-dried tissue with a blade grinder or other alternative to the TissueRuptor.
2.Transfer tissue to a screw-capped 2-ml polypropylene tube. Mince tissue to a fine powder in a TissueLyser LT through rapid movement of stainless-steel beads in the tube. Once a fine powder is achieved, remove and recycle stainless-steel beads and keep dried powdered tissue in the screw-capped tube at −80°C.
Extraction of secondary metabolites
3.Weigh out 10 mg (±1 mg) dried powdered tissue (leaves, stems, or roots; see steps 1 and 2) in a 2-ml microcentrifuge tube per sample. Prepare an additional empty tube as a blank for the extraction.
4.Add 400 µl LC-MS-grade water and 400 µl methanol and mix using a vortex.
5.Agitate samples for 1 hr at room temperature in a tube rotator and protect from light by covering sample holder with aluminum foil.
6.Add 800 µl chloroform and mix well using a vortex.
7.Centrifuge 3 min at maximum speed in a microcentrifuge to allow phase separation.
8.Recover top phase (epiphase), containing polar and semi-polar metabolites, and filter through a 2-mm syringe filter with a 0.45-µm nylon membrane using a 1-ml syringe.
9.Place filtered extract in a 1.5-ml microcentrifuge tube and analyze immediately or store at −20°C until analysis.
10.Recover lower organic layer (hypophase), containing non-polar metabolites, and discard middle layer, which contains precipitated proteins and other non-extracted compounds/macromolecules.
11.Transfer organic layer to a 1.5-ml microcentrifuge tube and dry under an N2 stream or using a vacuum centrifuge.
Analysis by LC-MS
12.Prior to analysis, centrifuge methanolic extract (epiphase; see steps 8 and 9) for 5 min at maximum speed to clean up sample.
13.Transfer an aliquot of clean extract to a conical glass insert mounted in a screw-capped 2-ml glass vial.
14.Spike extract with internal standard (0.2 mg/ml genistein in ethanol) so that the final concentration in the vial is 0.01 mg/ml and the spiked volume is <5% of the total volume to avoid dilution effects on the sample's metabolites.
15.Close vial with a cap with a PTFE/red silicone septum.
16.Use an injection volume of ≤5 µl for analysis in an LC-MS system with an electrospray source. To prevent degradation artifacts, keep sample at 6°C in the automated sampler.
17.Perform chromatographic separation of cassava's secondary metabolites using a linear gradient of acidified (0.01% to 0.1% formic acid) water (solvent A) and acetonitrile (solvent B) on a reverse-phase C18 analytical column operating at 30° to 35°C and a flow rate of 0.2 ml/min.
Quantification
18.For relative quantification, divide the area of each chemical feature's extracted ion chromatogram (EIC) (Ax) by the area of the internal standard's EIC (Ais) and the amount of analyte (Cx) calculated respective to the known amount of internal standard (Cis) added to the sample; this follows the classic formula of internal calibration: relative response factor (RRF) = [(Ax)(Cis)]/[(Ais)(Cx)].
19.For accurate quantification of candidate molecular features, perform external and internal calibration.
Basic Protocol 8: DETERMINATION OF CAROTENOIDS, CHLOROPHYLLS, AND TOCOPHEROL
Carotenoids represent the largest class of pigments found in nature and are present in bacteria, fungi, algae, higher plants, and animals. To date, over 700 carotenoid structures have been reported. Industrially, they are used across multiple commercial sectors as colorants, supplements, and bioactives. In plants, they act as potent antioxidants, ancillary photosynthetic pigments, and phytohormone precursors. An inherent problem associated with analysis of carotenoids is their poor ionization, which limits mass spectral determination. However, carotenoid pigments do have characteristic UV/Vis spectra due to the chromophore present in their chemical structure. This has been exploited through use of a photodiode array detector (PDA) coupled to on-line chromatographic separation. The method provided uses a routine ultra-performance liquid chromatography (UPLC) platform coupled to an on-line PDA.
The extraction method presented here is adapted from the classic Bligh and Dyer method (Bligh & Dyer, 1959). The method has been validated and extensively applied to detection and quantification of carotenoids in numerous crops (Drapal, Barros de Carvalho, et al., 2019; Drapal, Rossel, Heider, & Fraser, 2019; Price, Bhattacharjee, Lopez-Montes, & Fraser, 2018; Uluisik et al., 2016).
Additional Materials (also see Basic Protocol 7)
-
50 mM Tris buffer, pH 7.5 (Sigma, T1503)
-
Ethyl acetate (HPLC grade; Fisher Scientific, E/0906/17)
-
Mobile phase A: 50:50 (v/v) methanol (HPLC grade; Fisher Scientific, M/4056/17)/water (HPLC grade; Fisher Scientific, W/0106/17)
-
Mobile phase B: 75:25 (v/v) acetonitrile (HPLC grade; Fisher Scientific, 83640.320)/ethyl acetate
-
Standards:
- β-Carotene (Sigma-Aldrich, PHR1239; use also as reference standard for epoxy derivatives of β-carotene)
- Lutein (Sigma-Aldrich, 07168)
- Phytoene (purified by collecting HPLC fractions from Phycomyces blakesleeanus carB mutant extract; also available from Sigma-Aldrich, 78903)
- Violaxanthin (Sigma-Aldrich, 91904)
- Neoxanthin (Sigma-Aldrich, 72994)
- Chlorophyll a (Sigma-Aldrich, 96145)
- Chlorophyll b (Sigma-Aldrich, 00538)
- α-Tocopherol (Sigma-Aldrich, 47783)
-
UPLC-PDA system with automated sampler
-
Reverse-phase C18 analytical column (100 × 2.1 mm, i.d. 1.7 µm)
Sample preparation/grinding
1.Perform sample preparation/grinding on freeze-dried cassava tissue according to Basic Protocol 7, steps 1 and 2.
Extraction of secondary metabolites
2.Weigh out 10 mg (±1 mg) dried powdered leaf or stem tissue or 50 mg dried powdered root tissue in a 2-ml microcentrifuge tube. Prepare an additional empty tube as a blank for the extraction.
3.Add 400 µl methanol and 800 µl chloroform and mix using a vortex.
4.Add 400 µl of 50 mM Tris buffer (pH 7.5) and mix well using vortex.
5.Centrifuge 3 min at maximum speed in a microcentrifuge to allow phase separation.
6.Discard top and middle phases.
7.Recover lower organic layer (hypophase), containing the isoprenoids, and transfer to a 1.5-ml microcentrifuge tube.
8.Dry under an N2 stream or using a vacuum centrifuge.
Analysis by UPLC-PDA
9.Re-dissolve dried extract in 50 µl ethyl acetate and centrifuge 10 min at maximum speed in a microcentrifuge to clean up sample.
10.Transfer 30 µl clean extract to a conical glass insert mounted in a screw-capped 2-ml glass vial.
11.Use an injection volume of ≤3 µl for analysis in a UPLC-PDA system.
12.Perform chromatography in reverse-phase mode using a reverse-phase C18 analytical column and a linear gradient of mobile phase A (methanol/water, 50:50), and mobile phase B (acetonitrile/ethyl acetate, 75:25). Operate column at 30°C and a flow rate of 0.6 ml/min and keep automated sampler temperature at 6°C.
13.Perform detection using a UV/Vis scan ranging from 200 to 600 nm.
Quantification
14.Perform quantification by peak integration at fixed-wavelength chromatograms: 450 nm for chlorophylls, β-carotene, and xanthophylls (Fig. 2A) and 286 nm for α-tocopherol and phytoene (Fig. 2B).
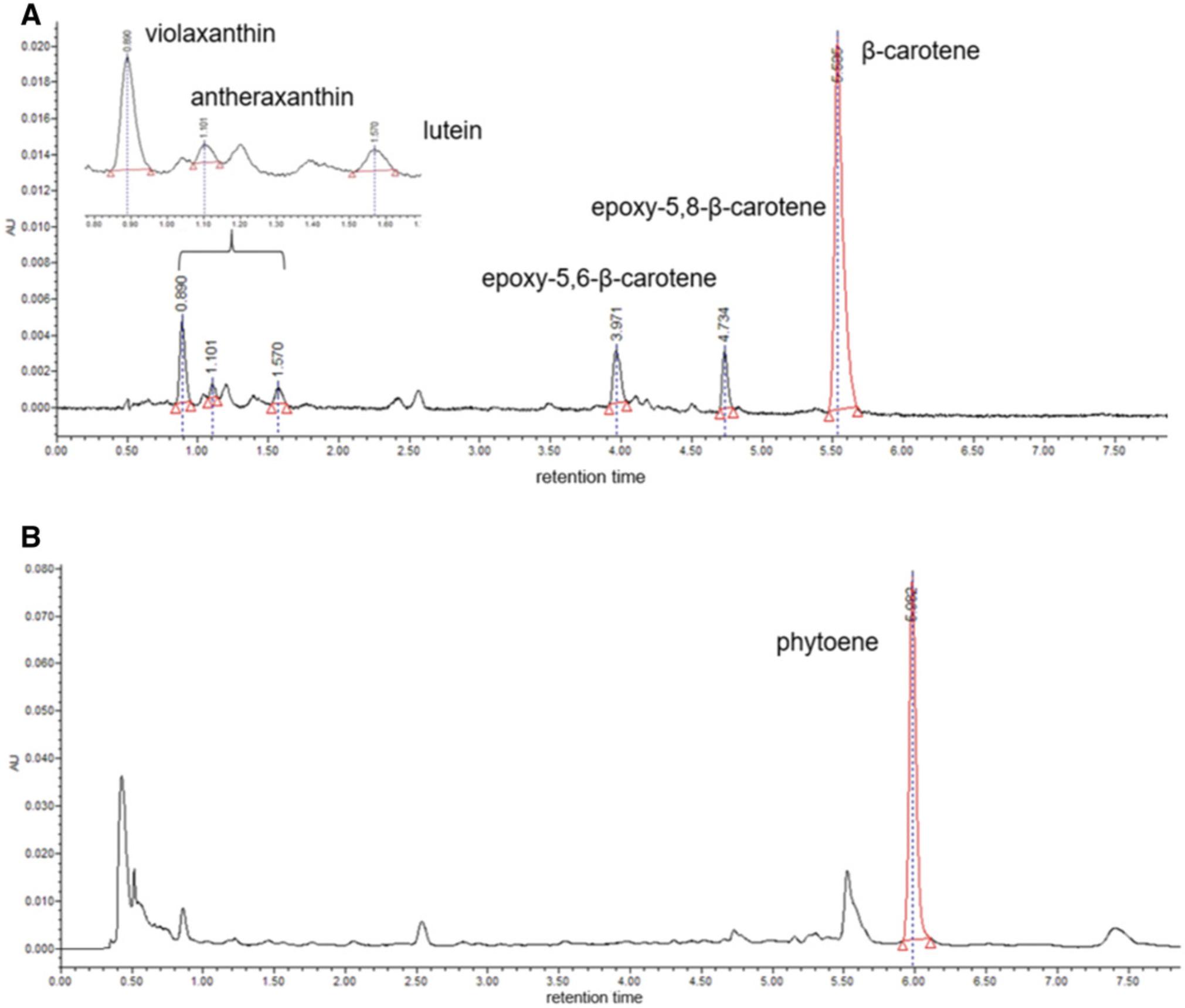
15.Perform external quantification by running external calibration curves of either commercial or purified standards.
Basic Protocol 9: DETERMINATION OF STARCH QUALITY
Whereas Basic Protocol 3 is useful for determining starch content in tissues, the protocol below allows assessment of the composition and structure of starch itself. These traits define properties of starch that influence its quality (e.g., viscosity), determining its end use.
Starch is the main metabolite that accumulates in cassava storage roots. It is composed of two glucose polymers consisting of linear chains linked by α-1,4-bonds that can be branched with other chains by α-1,6-bonds. Amylose is the minor glucan polymer, composing ≤30% of the starch and containing few branch points, whereas amylopectin is the major glucan polymer, is highly branched, and gives rise to the semi-crystalline structure of starch (Pfister & Zeeman, 2016). Glucan chains are naturally phosphorylated. The physicochemical properties of starch will influence the behavior of byproducts, such as the cooking quality of flour, or industrial applications of purified starch. We propose methods to assess the size of the starch granules, their amylose/amylopectin ratio, their chain-length distribution (CLD), and their phosphate content. The protocol presented here is optimized for cassava and is based on our previously described methods (Hostettler et al., 2011) and adapted from an existing protocol (Cledat, Battu, Mokrini, & Cardot, 2004).
Materials
-
Freeze-dried cassava storage-root powder (see Support Protocol 1)
-
Starch extraction buffer (see recipe)
-
Percoll cushion: 95% (v/v) Percoll (Sigma, P1644) and 5% (v/v) 0.5 M Tris·HCl, pH 8.0
-
0.5% (w/v) SDS
-
Ultrapure water (∼0.055 µS/cm; e.g., generated by Milli-Q Reference System)
-
80% (v/v) ethanol
-
Sterile ultrapure water
-
10% (v/v) Lugol solution (Sigma, 32922)
-
Debranching mix (see recipe; make fresh)
-
Dowex 1X8-200 (Dowex 1; 200 to 400 mesh, ion-exchange resin; Acros Organics, CAS 12627-85-9)
-
Dowex 50X2-100 (Dowex 50; 100 to 200 mesh, ion-exchange resin; Acros Organics, CAS 12612-37-2)
-
1 M sodium acetate (Sigma, CAS 127-09-03)
-
2 M hydrochloric acid (HCl; Roth, CAS 7647-01-0)
-
Liquid nitrogen
-
Eluent A: 100 mM NaOH (Sigma, CAS 1310-73-2)
-
Eluent B: 150 mM NaOH and 500 mM sodium acetate (Sigma, CAS 127-09-03)
-
1 M NaOH
-
5000 U/ml Antarctic phosphatase (New England Biolabs)
-
10× Antarctic phosphatase reaction buffer (New England Biolabs)
-
1 mM potassium phosphate monobasic (K2HPO4; Sigma, CAS 7778-77-0)
-
Malachite Green solution: one volume of 10% (w/v) ammonium molybdate tetrahydrate (Sigma, CAS 12054-85-2) in 4 M HCl and three volumes of 0.2% (w/v) Malachite Green (Sigma, CAS 569-64-2) in distilled water
-
Blender
-
Gauze
-
Funnel
-
70-µm nylon mesh
-
50-ml Falcon tubes
-
Elastic band
-
Standard tabletop centrifuge
-
13-ml round-bottom tubes
-
Analytical scale
-
96-well flat-bottom microtiter plates
-
Flow cytometer (Beckman Coulter, CytoFLEX S)
-
Polymethyl methacrylate sizing microbeads (refractive index, 1.49; diameters of 1, 4, 8, 20, and 30 µm; Sigma, MFCD00198073)
-
1.5-ml reaction tubes with screw caps
-
Vortex
-
37°C and 95°C Thermomixers (e.g., Eppendorf Thermomixer comfort)
-
Microtiter plate reader
-
37°C and 95°C water baths
-
Microcentrifuge (e.g., Eppendorf Centrifuge 5417 R)
-
Glass beakers
-
Magnetic stir bar and stir plate
-
Columns
-
5-ml scintillation vial
-
Freeze drier
-
1.5-ml microcentrifuge tubes
-
Vials suitable for autosampler of HPAEC-PAD system
-
High-performance anion-exchange chromatography with pulsed amperometric detection (HPAEC-PAD) system (ICS 5000 HPLC, Dionex) with autosampler and column (CarboPac® PA200 column, Dionex)
-
Multichannel pipet
Starch purification
1.Mix 5 ml freeze-dried cassava storage-root powder with 30 ml starch extraction buffer in a blender for 30 s.
2.Filter suspension first using gauze on top of a funnel to remove coarse pieces and then using 70-µm nylon mesh attached to the top of a 50-ml Falcon tube with an elastic band.
3.Centrifuge filtrate for 5 min at 3000 × g.
4.Resuspend pellet in 5 ml starch extraction buffer.
5.Load suspension onto a 4-ml Percoll cushion in a 13-ml round-bottom tube.
6.Centrifuge 5 min at 2500 × g.
7.Aspirate supernatant with a pipet.
8.Wash pellet in 5 ml of 0.5% SDS.
9.Centrifuge 3 min at 4500 × g.
10.Wash pellet twice with 10 ml ultrapure water.
11.Wash pellet once with 5 ml of 80% ethanol.
12.Dry starch at room temperature overnight.
Starch granule sizing
13.Resuspend ∼5 mg purified starch (see step 12) in 165 µl sterile ultrapure water.
14.Transfer 10 µl suspension to a well of a 96-well flat-bottom microtiter plate containing 190 µl sterile ultrapure water.
15.Measure 50,000 granules per sample using a flow cytometer (CytoFLEX S) configured with the following settings: forward scatter (FSC) gain, 20; threshold FSC, 6000; max 3000 events s−1; λ = 488 nm.
16.Use FSC values of different-sized polymethyl methacrylate sizing microbeads with a refractive index similar to that of moist starch granules to determine the correlation of the diameter to the FSC values of the sample.
Assessment of amylose content
17.Weigh out 5 mg purified starch (see step 12) in a 1.5-ml reaction tube with a screw cap.
18.Add 1 ml sterile ultrapure water and vortex thoroughly.
19.Incubate at 1000 rpm for 10 min at 95°C using a Thermomixer.
20.Immediately mix 5 µl gelatinized starch from step 19 with 200 µl of 10% Lugol solution in a 96-well flat-bottom microtiter plate.
21.Record OD550 and OD618 values on a microtiter plate reader.
22.Calculate percentage amylose content using the equation described for potato starch (Hovenkamp-Hermelink et al., 1988):
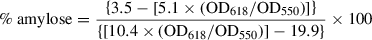
Assessment of chain-length distribution
23.Weigh out 0.2 mg purified starch (see step 12) in a 1.5-ml reaction tube with a screw cap.
24.Add 450 µl sterile ultrapure water and vortex thoroughly.
25.Incubate in a 95°C water bath for 15 min.
26.Cool sample to 20°C and spin briefly to remove condensed water and residual insoluble material from the tube cap.
27.Add 50 µl debranching mix to sample and mix well by vortexing.
28.Shake at 400 rpm for 3 hr at 37°C using a Thermomixer.
29.Stop reaction by heating the sample for 10 min in a 95°C water bath.
30.Spin down cell debris and denatured enzymes by centrifuging 10 min at 10,000 × g and reserve supernatant.
31.Prepare Dowex 1 and Dowex 50 ion-exchange columns to clean up the samples:
-
Weigh out each Dowex resin (1 g per sample) into separate glass beakers.
-
Wash resins in an excess of sterile ultrapure water. Let resins settle and then pour off water.
-
Charge Dowex 1 resin with 1 M sodium acetate and Dowex 50 resin with 2 M HCl using twice the resin volume. Stir for 30 min with a magnetic stir bar on a stir plate and then let resins settle.
-
Remove supernatant and wash resins with excess sterile ultrapure water four times or until the pH of the supernatant is neutral.
-
Prepare Dowex ion-exchange columns by stacking a Dowex 50 column above a Dowex 1 column. Place a 5-ml scintillation vial below the lower column.
We use in-house-made racks (Fig.3) so that the flow drops directly from one column to the next. A cotton pellet is placed at the bottom of each column to prevent the resin from flowing out. We can fit 96 samples in one rack. For an alternative system, see Hostettler et al. (2011).
- Resuspend charged resins in an equal volume of sterile ultrapure water, making a slurry. Mix each resin with a magnetic stir bar on a stir plate while pipetting it into the appropriate column.
The clean-up entails removal of neutral compounds and retention of charged compounds.
After the water has drained from the column, the resin bed volume should be 800 µl.
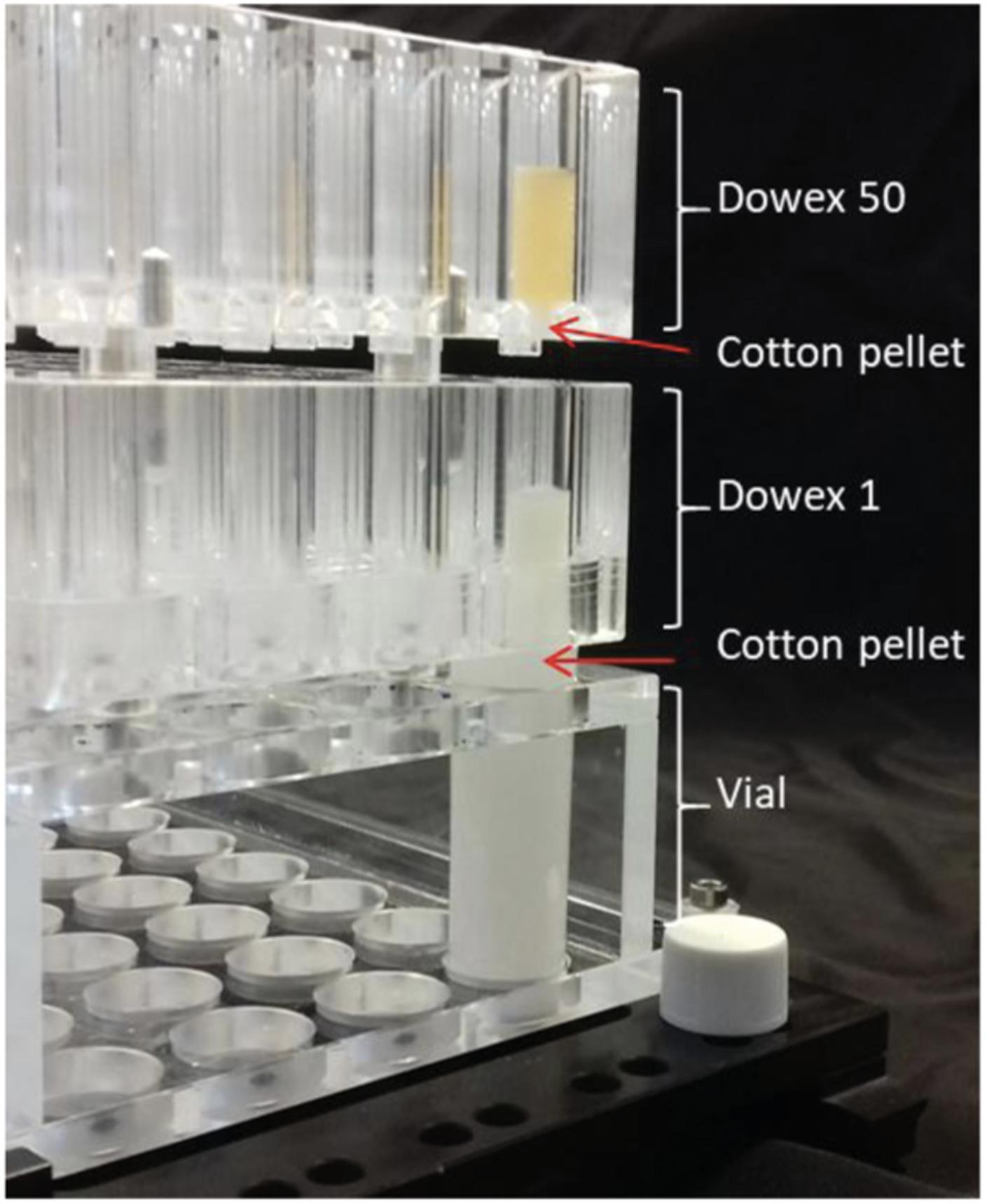

32.Apply supernatant of the centrifuged sample from step 30 to the Dowex 50 column. Let sample run into the resin.
33.Add 500 µl sterile ultrapure water and let it run into resin. Repeat this step every 5 min for a total of eight times to give a final elution volume of 4 ml.
34.Freeze sample in liquid nitrogen.
35.Freeze-dry sample until completely dry (usually ∼2 days).
36.Add 200 µl sterile ultrapure water to freeze-dried sample and dissolve well by vortexing.
37.Transfer dissolved sample to a 1.5-ml microcentrifuge tube and briefly spin down to remove any resin remnants.
38.Carefully transfer 100 µl supernatant to a vial suitable for autosampler of the HPAEC-PAD system. Perform HPAEC-PAD using HPAEC-PAD system with a column and the following standard program:
-
Starting conditions: 95% Eluent A and 5% Eluent B.
-
Gradient 0-13 min: linear to 60% Eluent A and 40% Eluent B.
-
Gradient 13-50 min: linear to 15% Eluent A and 85% Eluent B.
-
Re-equilibrating 50-70 min: 95% Eluent A and 5% Eluent B.
For HPAEC-PAD, we use an ICS 5000 HPLC (Dionex) system equipped with an electrochemical detector and gold electrode. We recommend using a CarboPac® PA200 column (Dionex), which is designed for separation of neutral oligosaccharides. Separation is achieved at a high pH, such that the hydroxyl groups are ionized. Longer glucans bind more tightly to the column. The chains are eluted with a NaOH/sodium acetate gradient.
We normally inject 10 µl for analysis.
39.Determine and sum individual peak areas. Express each peak area as a percentage of the total area of all peaks and plot values accordingly (Fig. 4).
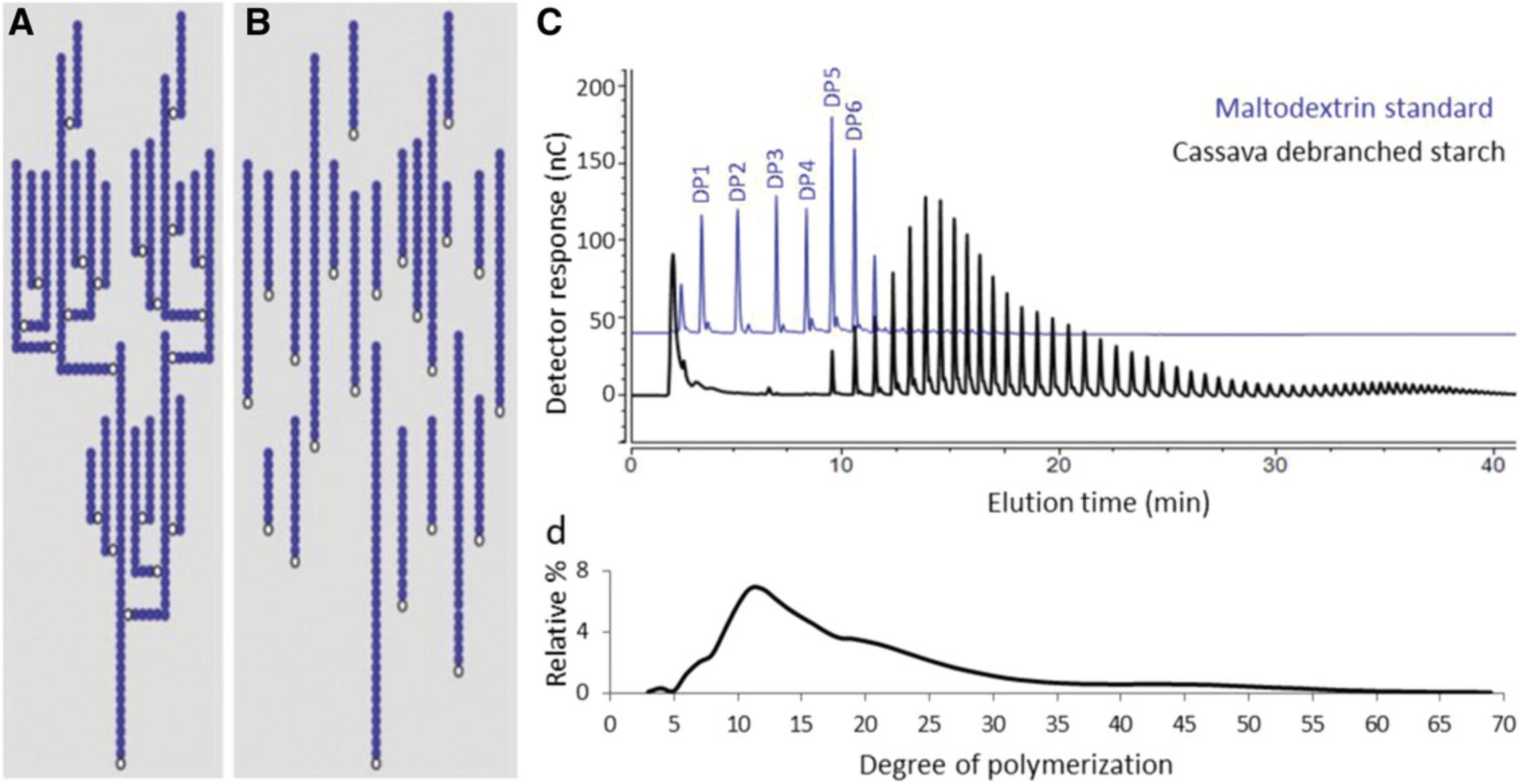
Determination of phosphate content
40.Weigh out 5 mg purified starch (see step 12) in a 1.5-ml reaction tube and add 50 µl ultrapure water. Prepare triplicates and also a blank sample without starch.
41.Add 50 µl of 2 M HCl and acid-hydrolyze for 2 hr in a 95°C water bath.
42.Cool samples to room temperature and neutralize with ∼100 µl of 1 M NaOH.
43.Centrifuge tubes for 5 min at 10,000 × g , room temperature. Transfer 50 µl hydrolyzed supernatant into a new 1.5-ml reaction tube and add 3 µl of Antarctic phosphatase and 5.3 µl of 10× Antarctic phosphatase reaction buffer. Mix by pipetting up and down.
44.Incubate for 2 hr in a 37°C water bath.
45.Transfer 30 µl dephosphorylated sample into a 96-well flat-bottom microtiter plate and add 20 µl ultrapure water.
46.Prepare a calibration curve using a dilution series of 0 to 10 nmol K2HPO4 using 1 mM K2HPO4 stock solution.
47.Using a multichannel pipet, add 125 µl Malachite Green solution to each well.
48.Incubate 10 min at room temperature and read absorbance at 660 nm with a microtiter plate reader.
49.Plot absorbance of the standards against the amount of phosphate. Determine linear range and calculate slope of the standard curve. Then, calculate phosphate content of the starch samples, taking into account dilutions made during the procedure.
REAGENTS AND SOLUTIONS
Debranching mix
- 100 mM sodium acetate/acetic acid, pH 4.8 (filter-sterilized; Sigma, CAS 127-09-03)
- 0.8 U/ml isoamylase (stock 1000 U/ml from Pseudomonas sp.; Megazyme, CAS 9067-73-6)
- 20 U/ml pullulanase (stock 700 U/ml from Klebsiella planticola ; Megazyme, CAS 9075-68-7)
- Prepare fresh immediately before use
Glucose assay mix
- For a final volume of 200 µl per well:
- 100 µl 100 mM 2-(4-[2-hydroxyethyl]-1-piperazinyl)-ethanesulfonic acid-KOH (HEPES-KOH), pH 7.5
- (50 mM final), with 2 mM magnesium chloride (MgCl2; 1 mM final)
- 20 µl 10 mM ATP (Roche, 10127523001; 1 mM final)
- 20 µl 10 mM nicotinamide adenine dinucleotide (NAD+, free acid, grade II; Roche, 10127981001; 1 mM final)
- 0.3 µl 1500 U/ml hexokinase (stock 1500 U/ml from Saccharomyces cerevisiae ; Roche, 11426362001; 2.25 U/ml final)
- X µl deionized H2O (depending on sample volume or glucose-standard volume)
- Store ≤7 days at 4°C
Protein extraction buffer
- 200 mM sodium chloride (NaCl; pharma grade, AppliChem, 141659.1214)
- 1 mM EDTA (EDTA disodium salt 2-hydrate, or Na-EDTA; p.a.; AppliChem, 131669.1211, CAS 6381-92-6), pH 7.8
- 0.2% (v/v) Triton® X-100 (Sigma-Aldrich, T9284-1l, CAS 9002-93-1)
- 100 mM Tris·HCl (Tris ultrapure, AppliChem, A1086, CAS 77-86-1), pH 7.8
- 1% (v/v) β-mercaptoethanol (AppliChem, A1108, CAS 60-24-2)
- 1 ×cOmpleteTM Protease Inhibitor Cocktail (to prepare a 25× stock, use one tablet per 2 ml buffer, e.g., PBS or extraction buffer without β-mercaptoethanol; Roche/Sigma-Aldrich, 11697498001)
- Store ≤7 days at 4°C
- Immediately before use, add the following:
Starch digestion mix (for Basic Protocol 3)
- 200 mM sodium acetate (C2H3NaO2; Sigma-Aldrich, 32319-1KG-R, CAS 127-09-3), pH 4.8 (adjust pH with NaOH)
- 6.3 U/ml amyloglucosidase (from Aspergillus niger; Roche/Merck, 10102857001)
- 50 U/ml ɑ-amylase (Sigma-Aldrich, 10070)
- Prepare fresh immediately before use
Starch digestion mix (for Alternate Protocol)
- 220 mM sodium acetate/acetic acid pH 4.8 (Sigma, CAS 127-09-03)
- 6.3 U/ml amyloglucosidase (stock 10 mg/ml from Aspergillus niger , Roche, 10102857001)
- 50 U/ml ɑ-amylase (stock 10 mg/ml; Roche, 10102814001)
- Prepare fresh immediately before use
Starch extraction buffer
- 50 mM Tris·HCl (Tris ultrapure, AppliChem, A1086, CAS 77-86-1), pH 8
- 0.2 mM EDTA (EDTA disodium salt 2-hydrate, or Na-EDTA; p.a.; AppliChem, 131669.1211, CAS 6381-92-6), pH 8
- 0.5% (v/v) Triton® X-100 (Sigma-Aldrich, T9284-1l, CAS 9002-93-1)
- Store ≤7 days at 4°C
COMMENTARY
Background Information
Since the conceptual framework of metabolite analysis as one of the “omics” approaches was laid at the end of the last century (Oliver, Winson, Kell, & Baganz, 1998), metabolomics has been a constant evolving field due to improvements in materials, machines, and techniques. Even though a large number of metabolites still remain to be identified, the contributions of metabolomics to different areas of plant science range from fundamental knowledge on genetic and biochemical mechanisms to practical applications in crop quality improvement and food safety assessment.
Currently, the chemical diversity of such molecules makes it impossible to cover all metabolite classes present in plant cells using a single technical approach. In order to obtain an overview of the different metabolite classes, we highly recommend joining forces with specialized laboratories with experience in using different protocols. In some cases, these laboratories have unique expertise in several aspects of metabolome analysis; however, central technologies can be carried out by more than one laboratory. On the one hand, this allows increased throughput of time-demanding measurements, and on the other hand, it allows duplicate measurements to further improve the value and quality of the data generated.
It is also important to mention that the quality of our measurements of cassava metabolites was achieved by adopting quality-control (QC) approaches, namely recovery and recombination. In the former (Arrivault et al., 2009), a defined amount of a known standard is added to frozen plant material before extraction, and how much of this defined amount can be recovered following extraction, handling, chromatography, and other processing is assessed. Ideally, the amount of added standard should be similar to or at most two- to three-fold above that in the extract. This is because time-dependent loss or loss due to reaction with components that are present in small amounts in the extract may not be detected if the standard is added in large excess. The reproducibility of recovery in technical or biological replicates also provides a good test of the precision of the sample handling and analytic procedures. In recombination, a standard plant extract (e.g., from unstressed Arabidopsis leaves) is mixed with cassava extract prior to extraction, and two questions are asked: firstly, do peaks co-elute between samples, and secondly, is the mixed sample equal to the sum of the parts from which it is derived? Recovery and recombination controls can also be performed after extraction and prior to chromatography to test for loss post-extraction but during analysis. This is a good first step in troubleshooting in the case that poor results are obtained when recovery or recombination is performed with unextracted material because it demarcates if the major problem arises during extraction or in the procedures used to measure the metabolite.
Due to logistic limitations in shipping field-grown cassava samples, freeze-dried material was established as the standard for some of the analysis. Therefore, the effects of freeze-drying on detection of metabolites has also been analyzed. In this test, frozen cassava material derived from identical pools is split into two aliquots, with one of them analyzed fresh and the other analyzed after freeze-drying.
The protocols presented in this article allow relative and/or absolute quantification of metabolites comprised by different pathways, from the core central and highly abundant metabolites present in plant and non-plant species, such as sugars and organic acids, to more specialized (also known as secondary; Alseekh & Fernie, 2018) metabolites present in cassava, such as linamarin. The modifications and optimizations of metabolomics techniques to apply to cassava tissues, as presented in this article, have been helping to and will continue to expand our knowledge about such an interesting and essential crop.
Critical Parameters
In plant metabolomics experiments, environmental, biological, and/or technical noise is unavoidable. Several factors can contribute to variability across samples, and therefore, appropriate experimental design should be considered a priori and carefully thought through. For detailed metabolic analysis, different plant tissues can be harvested at the desired developmental stage and/or time points. We recommend constraining the sampling time to the minimum possible to minimize diurnal deviation in metabolite levels (Fernie et al., 2011). Samples from each tissue should be snap-frozen in liquid nitrogen immediately after sampling and freeze-dried for the appropriate time (Support Protocol 1) according to the characteristics of each tissue. Some metabolites can have fast half-lives in vivo (<1 s for Calvin-Benson cycle intermediates, ADP glucose, and sucrose 6-phosphate; Arrivault et al., 2009; <1 min for many glycolytic metabolites); it is therefore important to avoid even transient shading of leaf material before quenching. For bulky tissues, it must be kept in mind that although the surface freezes quickly, cooling to the center is slow. Here, it is advisable to make thin slices and to allow each slice to drop immediately into liquid nitrogen (Geigenberger, Fernie, Gibon, Christ, & Stitt, 2000). Collection of slices and freezing after 30 to 60 s is not advisable because oxygen levels can be low inside bulky tissues and then rise rapidly after slicing, leading to changes in the levels of metabolites that are linked to respiratory ATP formation. For other metabolites, like sugars, amino acids, organic acids, and specialized metabolites as well as ions, the turnover times are slower and less rigorous, so easier harvesting methods can be considered.
For large sampling sets, we recommend randomized block design and use of QC samples every 20 to 25 samples to correct for batch-to-batch variation, especially when using a mass spectrometer as the detection system. QC samples are typically a pool of the samples included in the batch or can be a reference material. The QC samples are extracted and processed as the samples are.
Samples should be well ground to a fine powder (Support Protocol 1 and Basic Protocol 2) in order to have homogeneous material for each analysis. During the grinding procedure, the material and all utensils (e.g., tubes, Retsch balls, mortar, pestle) must be cooled using liquid nitrogen or dry ice to avoid heating of the material and degradation of metabolites.
After grinding, we recommend a second round of freeze-drying to avoid the presence of water during the extraction procedure. Once extracted, the chemical stability of metabolites in solution is critical. Analysis of metabolites immediately after extraction is recommended. However, if transport or storage of extracts is involved in the process, then drying the extracts and storage at low temperature (−20°C or −80°C) aid in preventing chemical degradation of compounds. Extracts can be stored at −80°C for ≤3 months. A longer storage time has not yet been systematically investigated.
Troubleshooting
In the case that large variation between replicates is observed in Basic Protocol 7, the tissue might not have been ground adequately, or the pipetting may not have been consistent. If chromatographic peaks are too broad/have a flat top, the sample is too concentrated and needs dilution to the appropriate level. If metabolites are detected in the blank extraction, contamination of solvents is possible, which requires re-extraction with new solvents.
Validation recovery experiments are conducted to account for extraction losses and matrix effects. Creation of standard curves and recovery experiments are described in Support Protocol 4.
For protocols in which standard curves are used, the values for the samples should fit within the linear range of the standard curve. If the values for the samples are higher than the linear range of the standard curve, either lower the volume of the samples being analyzed or dilute the extract with the appropriate solution according to the protocol. In general, the extraction buffer or the buffer in which the samples were resuspended should be used for the dilution. For Basic Protocol 6, sometimes, protein standard curves are nonlinear with increasing amounts of the standard protein. In that case, extend the time of incubation (of samples and standards) to up to 30 min, instead of the routinely used 15 min.
Understanding Results
The combination of all protocols described allows quantification of approximately over 100 metabolites, including sugars, organic acids, amino acids, phosphorylated intermediates, minerals, starch, total protein, and secondary metabolites, depending on the tissue type.
Time Considerations
The necessary time varies greatly according to the protocol used; personal experience; and resources available in the laboratory, such as the number of centrifuges and vacuum concentrators. Here, we give estimated times for each procedure. We highly recommend use of electronic pipets when possible.
In Basic Protocol 1, around 60 samples can easily be extracted per day, and the drying step can be done overnight. For the derivatization, we recommend preparation of approximately 30 samples per day, including standards, as each sample needs to be run in splitless and split modes, consuming double the time.
The time necessary for harvest and preparation of samples in Support Protocol 1 can vary greatly according to the size of the experiment and the number and type of samples to be collected. The harvest time constraints mentioned in the Critical Parameters section also need to be considered. As an example, 30 leaf samples can be harvested and further prepared within 8 days.
For the preparation of standard compounds described in Support Protocol 2, once the stock solution for each metabolite is ready, both mixes can be prepared within 2 days, whereas Support Protocol 3 can be performed in 1 day.
For Basic Protocol 2, extraction of phosphorylated metabolites and carbonic acids from a small set of samples (up to 12) takes ∼120 min. The machine runtime of this sample set is ∼30 hr (∼2 hr per sample). It is advised to freshly extract the material prior to measurement, as some of the extracted metabolites break down quickly. Extracted samples can be stored frozen at −80°C for approximately 3 to 4 days.
In Support Protocol 4, initial generation of the standard mix and standard dilutions takes ∼3 hr. However, the undiluted standard mix can be stored at −20°C for several months, and subsequent dilutions from the mix can be done within minutes. Spike-in experiments consist of metabolite extraction, measurement, and data evaluation. Extraction of phosphorylated metabolites and carbonic acids from a small set of samples (up to 12) takes ∼120 min. The machine runtime of this sample set is approximately 30 hr (∼2 hr per sample). It is advised to freshly extract the material prior to measurement, as some of the extracted metabolites break down quickly. Extracted samples can be stored frozen for approximately 3 to 4 days. Data evaluation for this sample set takes ∼1 hr.
For Basic Protocol 3, up to 72 samples can easily be extracted/processed within 1 day, e.g., when performing the three different extraction procedures for a batch of 24 samples each and the following digestion of the starch for the 72 samples together. The determination can be done within 4 hr on the next day. For the Alternate Protocol, the full procedure for 96 samples takes 2 days.
Basic Protocol 4 can be done for a batch of 24 samples within 1 hr. Basic Protocol 5 takes ∼3 hr for nine samples plus a blank. Depending on the rotor (in our case, it has space for 10 buckets) of the microwave, more samples could be processed in the same time. Basic Protocol 6, including the Bradford assay, can be done within 8 hr for 48 samples.
The extraction process in Basic Protocol 7 for a standard sample set of 24 samples takes ∼90 min, followed by 60 min of sample preparation for the LC/MS. Depending on the column and method used, one sample can be analyzed in 10 to 30 min. The data analysis depends on the processing capacity of the computer, and a set of approximately 300 samples takes 1 week to process for relative quantification.
In Basic Protocol 8, extraction, including drying of extracts, takes ∼60 min for a standard sample set of 24 samples. The analysis by UPLC-PDA for a standard sample set includes ∼30 min of preparation of the sample set and ∼10 min of chromatographic separation per sample. See Basic Protocol 7 for timings of the data analysis.
In Basic Protocol 9, the necessary time varies for each step: for starch purification, approximately 30 samples per day can be processed; for starch granule sizing, the full procedure for 96 samples takes 3 hr; for amylose content assessment, the full procedure for 96 samples takes 1 day; for analysis of CLD, the full procedure for 96 samples takes 2 weeks; and for phosphate content determination, the full procedure for 96 samples takes 2 days.
Acknowledgments
This research was supported by the Bill and Melinda Gates Foundation (OPP1113365; “CASS — Metabolic engineering of carbon pathways to enhance yield of root and tuber crops”).
Funding from the Bill and Melinda Gates Foundation (OPP1058938) through the Natural Resources Institute at the University of Greenwich and funding from the CGIAR Research Program on Roots, Tubers and Bananas (RTB) supported by CGIAR donors are also acknowledged.
Literature Cited
- Alseekh, S., & Fernie, A. R. (2018). Metabolomics 20 years on: What have we learned and what hurdles remain? Plant Journal , 94, 933–942. doi: 10.1111/tpj.13950.
- Arrivault, S., Guenther, M., Ivakov, A., Feil, R., Vosloh, D., van Dongen, J. T., … Stitt, M. (2009). Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant Journal , 59, 826–839. doi: 10.1111/j.1365-313X.2009.03902.x.
- Bjarnholt, N., Neilson, E. H. J., Crocoll, C., Jorgensen, K., Motawia, M. S., Olsen, C. E., … Moller, B. L. (2018). Glutathione transferases catalyze recycling of auto-toxic cyanogenic glucosides in sorghum. Plant Journal , 94, 1109–1125. doi: 10.1111/tpj.13923.
- Bligh, E. G., & Dyer, W. J. (1959). A rapid method of total lipid extraction and purification. Canadian Journal of Biochemistry and Physiology , 37, 911–917. doi: 10.1139/y59-099.
- Bradford, M. M. (1976). Rapid and sensitive method for quantitation of microgram quantities of protein utilizing principle of protein-dye binding. Analytical Biochemistry , 72, 248–254. doi: 10.1016/0003-2697(76)90527-3.
- Britton, G., Liaaen-Jensen, S., & Pfander, H. (1994). Carotenoids, volume 1B: Spectroscopy. Basel: Birkhäuser.
- Britton, G., Pfander, H., & Liaaen-Jensen, S. (2004). Carotenoids handbook. Basel; Boston: Birkhäuser Verlag. c2004. 1v.
- Chandrasekara, A., & Josheph Kumar, T. (2016). Roots and tuber crops as functional foods: A review on phytochemical constituents and their potential health benefits. International Journal of Food Science , 2016, 3631647. doi: 10.1155/2016/3631647.
- Chetty, C. C., Rossin, C. B., Gruissem, W., Vanderschuren, H., & Rey, M. E. (2013). Empowering biotechnology in southern Africa: Establishment of a robust transformation platform for the production of transgenic industry-preferred cassava. New Biotechnology , 30, 136–143. doi: 10.1016/j.nbt.2012.04.006.
- Cledat, D., Battu, S., Mokrini, R., & Cardot, P. J. (2004). Rice starch granule characterization by flow cytometry scattering techniques hyphenated with sedimentation field-flow fractionation. Journal of Chromatography A , 1049, 131–138. doi: 10.1016/j.chroma.2004.08.003.
- De Souza, A. P., Massenburg, L. N., Jaiswal, D., Cheng, S., Shekar, R., & Long, S. P. (2017). Rooting for cassava: Insights into photosynthesis and associated physiology as a route to improve yield potential. New Phytologist , 213, 50–65. doi: 10.1111/nph.14250.
- Drapal, M., Barros de Carvalho, E., Ovalle Rivera, T. M., Becerra Lopez-Lavalle, L. A., & Fraser, P. D. (2019). Capturing biochemical diversity in Cassava (Manihot esculenta Crantz) through the application of metabolite profiling. Journal of Agricultural and Food Chemistry , 67, 986–993. doi: 10.1021/acs.jafc.8b04769.
- Drapal, M., Rossel, G., Heider, B., & Fraser, P. D. (2019). Metabolic diversity in sweet potato (Ipomoea batatas, Lam.) leaves and storage roots. Horticulture Research , 6, 2. doi: 10.1038/s41438-018-0075-5.
- Fernie, A. R., Aharoni, A., Willmitzer, L., Stitt, M., Tohge, T., Kopka, J., … DeLuca, V. (2011). Recommendations for reporting metabolite data. Plant Cell , 23, 2477–2482. doi: 10.1105/tpc.111.086272.
- Fernie, A. R., & Pichersky, E. (2015). Focus issue on metabolism: Metabolites, metabolites everywhere. Plant Physiology , 169, 1421–1423. doi: 10.1104/pp.15.01499.
- Fernie, A. R., Trethewey, R. N., Krotzky, A. J., & Willmitzer, L. (2004). Metabolite profiling: From diagnostics to systems biology. Nature Reviews Molecular Cell Biology , 5, 763–769. doi: 10.1038/nrm1451.
- Fraser, P. D., Pinto, M. E., Holloway, D. E., & Bramley, P. M. (2000). Technical advance: Application of high-performance liquid chromatography with photodiode array detection to the metabolic profiling of plant isoprenoids. Plant Journal , 24, 551–558. doi: 10.1046/j.1365-313x.2000.00896.x.
- Geigenberger, P., Fernie, A. R., Gibon, Y., Christ, M., & Stitt, M. (2000). Metabolic activity decreases as an adaptive response to low internal oxygen in growing potato tubers. Biological Chemistry , 381, 723–740. doi: 10.1515/BC.2000.093.
- Hall, R. D., & Hardy, N. W. (2012). Practical applications of metabolomics in plant biology. Methods in Molecular Biology , 860, 1–10. doi: 10.1007/978-1-61779-594-7_1.
- Hofmann, J., Bornke, F., Schmiedl, A., Kleine, T., & Sonnewald, U. (2011). Detecting functional groups of Arabidopsis mutants by metabolic profiling and evaluation of pleiotropic responses. Frontiers in Plant Science , 2, 82. doi: 10.3389/fpls.2011.00082.
- Hostettler, C., Kolling, K., Santelia, D., Streb, S., Kotting, O., & Zeeman, S. C. (2011). Analysis of starch metabolism in chloroplasts. Methods in Molecular Biology , 775, 387–410. doi: 10.1007/978-1-61779-237-3_21.
- Hovenkamp-Hermelink, J. H. M., De Vries, J. N., Adamse, P., Jacobsen, E., Witholt, B., & Feenstra, W. J. (1988). Rapid estimation of the amylose/amylopectin ratio in small amounts of tuber and leaf tissue of the potato. Potato Research , 31, 241–246. doi: 10.1007/BF02365532.
- Howeler, R., Lutaladio, N., & Thomas, G. (2013). Save and grow: Cassava. A guide to sustainable production intensification. Rome, Italy: Food and Agriculture Organization of the United Nations.
- Klaumann, S., Nickolaus, S. D., Furst, S. H., Starck, S., Schneider, S., Neuhaus, H. E., & Trentmann, O. (2011). The tonoplast copper transporter COPT5 acts as an exporter and is required for interorgan allocation of copper in Arabidopsis thaliana. New Phytologist , 192, 393–404. doi: 10.1111/j.1469-8137.2011.03798.x.
- Lei, Z., Huhman, D. V., & Sumner, L. W. (2011). Mass spectrometry strategies in metabolomics. Journal of Biological Chemistry , 286, 25435–25442. doi: 10.1074/jbc.R111.238691.
- Lisec, J., Schauer, N., Kopka, J., Willmitzer, L., & Fernie, A. R. (2006). Gas chromatography mass spectrometry-based metabolite profiling in plants. Nature Protocols , 1, 387–396. doi: 10.1038/nprot.2006.59.
- Luedemann, A., Strassburg, K., Erban, A., & Kopka, J. (2008). TagFinder for the quantitative analysis of gas chromatography - mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics , 24, 732–737. doi: 10.1093/bioinformatics/btn023.
- Luedemann, A., von Malotky, L., Erban, A., & Kopka, J. (2012). TagFinder: Preprocessing software for the fingerprinting and the profiling of gas chromatography-mass spectrometry based metabolome analyses. Methods in Molecular Biology , 860, 255–286. doi: 10.1007/978-1-61779-594-7_16.
- Montagnac, J. A., Davis, C. R., & Tanumihardjo, S. A. (2009). Nutritional value of Cassava for use as a staple food and recent advances for improvement. Comprehensive Reviews in Food Science and Food Safety , 8, 181–194. doi: 10.1111/j.1541-4337.2009.00077.x.
- Oliver, S. G., Winson, M. K., Kell, D. B., & Baganz, F. (1998). Systematic functional analysis of the yeast genome. Trends in Biotechnology , 16, 373–378. doi: 10.1016/S0167-7799(98)01214-1.
- Pfister, B., & Zeeman, S. C. (2016). Formation of starch in plant cells. Cellular and Molecular Life Sciences , 73, 2781–2807. doi: 10.1007/s00018-016-2250-x.
- Picmanova, M., Neilson, E. H., Motawia, M. S., Olsen, C. E., Agerbirk, N., Gray, C. J., … Bjarnholt, N. (2015). A recycling pathway for cyanogenic glycosides evidenced by the comparative metabolic profiling in three cyanogenic plant species. Biochemical Journal , 469, 375–389. doi: 10.1042/BJ20150390.
- Price, E. J., Bhattacharjee, R., Lopez-Montes, A., & Fraser, P. D. (2018). Carotenoid profiling of yams: Clarity, comparisons and diversity. Food Chemistry , 259, 130–138. doi: 10.1016/j.foodchem.2018.03.066.
- Uluisik, S., Chapman, N. H., Smith, R., Poole, M., Adams, G., Gillis, R. B., … Seymour, G. B. (2016). Genetic improvement of tomato by targeted control of fruit softening. Nature Biotechnology , 34, 950–952. doi: 10.1038/nbt.3602.
Citing Literature
Number of times cited according to CrossRef: 16
- Sindy Gutschker, David Ruescher, Ismail Y. Rabbi, Laise Rosado‐Souza, Benjamin Pommerrenig, Markus Pauly, Stefan Robertz, Anna M. Doorn, Armin Schlereth, H. Ekkehard Neuhaus, Alisdair R. Fernie, Stephan Reinert, Uwe Sonnewald, Wolfgang Zierer, Carbon usage in yellow‐fleshed Manihot esculenta storage roots shifts from starch biosynthesis to cell wall and raffinose biosynthesis via the myo‐inositol pathway, The Plant Journal, 10.1111/tpj.16909, 119 , 4, (2045-2062), (2024).
- David Rüscher, Viktoriya V Vasina, Jan Knoblauch, Leo Bellin, Benjamin Pommerrenig, Saleh Alseekh, Alisdair R Fernie, H Ekkehard Neuhaus, Michael Knoblauch, Uwe Sonnewald, Wolfgang Zierer, Symplasmic phloem loading and subcellular transport in storage roots are key factors for carbon allocation in cassava, Plant Physiology, 10.1093/plphys/kiae298, 196 , 2, (1322-1339), (2024).
- Francisco Bruno S. Freire, Eva G. Morais, Danilo M. Daloso, Toward the apoplast metabolome: Establishing a leaf apoplast collection approach suitable for metabolomics analysis, Plant Physiology and Biochemistry, 10.1016/j.plaphy.2024.109080, 215 , (109080), (2024).
- Shijia Wang, Ruimei Li, Yangjiao Zhou, Alisdair R. Fernie, Zhongping Ding, Qin Zhou, Yannian Che, Yuan Yao, Jiao Liu, Yajie Wang, Xinwen Hu, Jianchun Guo, Integrated Characterization of Cassava (Manihot esculenta) Pectin Methylesterase (MePME) Genes to Filter Candidate Gene Responses to Multiple Abiotic Stresses, Plants, 10.3390/plants12132529, 12 , 13, (2529), (2023).
- Christian E. Lamm, Ismail Y. Rabbi, David Barbosa Medeiros, Laise Rosado‐Souza, Benjamin Pommerrenig, Ismail Dahmani, David Rüscher, Jörg Hofmann, Anna M. Doorn, Armin Schlereth, H. Ekkehard Neuhaus, Alisdair R. Fernie, Uwe Sonnewald, Wolfgang Zierer, Efficient sugar utilization and transition from oxidative to substrate‐level phosphorylation in high starch storage roots of African cassava genotypes, The Plant Journal, 10.1111/tpj.16357, 116 , 1, (38-57), (2023).
- Laura Perez-Fons, Tatiana Maria Ovalle, Margit Drapal, Maria Alejandra Ospina, Anestis Gkanogiannis, Adriana Bohorquez-Chaux, Luis Augusto Becerra Lopez-Lavalle, Paul David Fraser, Integrated genetic and metabolic characterization of Latin American cassava ( Manihot esculenta ) germplasm , Plant Physiology, 10.1093/plphys/kiad269, 192 , 4, (2672-2686), (2023).
- Nuti Hutasingh, Hathaichanok Chuntakaruk, Apinya Tubtimrattana, Yanisa Ketngamkum, Putthamas Pewlong, Narumon Phaonakrop, Sittiruk Roytrakul, Thanyada Rungrotmongkol, Atchara Paemanee, Nat Tansrisawad, Ubonrat Siripatrawan, Supaart Sirikantaramas, Metabolite profiling and identification of novel umami compounds in the chaya leaves of two species using multiplatform metabolomics, Food Chemistry, 10.1016/j.foodchem.2022.134564, 404 , (134564), (2023).
- Mercedes Thieme, Anton Hochmuth, Theresa Elisabeth Ilse, Jose A. Cuesta-Seijo, Szymon Stoma, Roger Meier, Simon Flyvbjerg Nørrelykke, Pai Rosager Pedas, Ilka Braumann, Samuel C. Zeeman, Detecting variation in starch granule size and morphology by high-throughput microscopy and flow cytometry, Carbohydrate Polymers, 10.1016/j.carbpol.2022.120169, 299 , (120169), (2023).
- Anna Vittoria Carluccio, Laure C. David, Joelle Claußen, Marco Sulley, Seun Raheemat Adeoti, Toyin Abdulsalam, Stefan Gerth, Samuel C. Zeeman, Andreas Gisel, Livia Stavolone, Set up from the beginning: The origin and early development of cassava storage roots, Plant, Cell & EnvironmentPlant, Cell & EnvironmentPlant, Cell & Environment, 10.1111/pce.14300, 45 , 6, (1779-1795), (2022).
- Margit Drapal, Laura Perez-Fons, Elliott J. Price, Delphine Amah, Ranjana Bhattacharjee, Bettina Heider, Mathieu Rouard, Rony Swennen, Luis Augusto Becerra Lopez-Lavalle, Paul D. Fraser, Datasets from harmonised metabolic phenotyping of root, tuber and banana crop, Data in Brief, 10.1016/j.dib.2022.108041, 42 , (108041), (2022).
- Dhika Amanda, Felix P. Frey, Ulla Neumann, Marine Przybyl, Jan Šimura, Youjun Zhang, Zongliang Chen, Andrea Gallavotti, Alisdair R. Fernie, Karin Ljung, Iván F. Acosta, Auxin boosts energy generation pathways to fuel pollen maturation in barley, Current Biology, 10.1016/j.cub.2022.02.073, 32 , 8, (1798-1811.e8), (2022).
- Alex C. Ogbonna, Punna Ramu, Williams Esuma, Leah Nandudu, Nicolas Morales, Adrian Powell, Robert Kawuki, Guillaume Bauchet, Jean-Luc Jannink, Lukas A. Mueller, A population based expression atlas provides insights into disease resistance and other physiological traits in cassava (Manihot esculenta Crantz), Scientific Reports, 10.1038/s41598-021-02794-y, 11 , 1, (2021).
- Hirofumi Ishihara, Thiago A. Moraes, Stéphanie Arrivault, Mark Stitt, Assessing Protein Synthesis and Degradation Rates in Arabidopsis thaliana Using Amino Acid Analysis, Current Protocols, 10.1002/cpz1.114, 1 , 5, (2021).
- Uwe Sonnewald, Alisdair R. Fernie, Wilhelm Gruissem, Pascal Schläpfer, Ravi B. Anjanappa, Shu‐Heng Chang, Frank Ludewig, Uwe Rascher, Onno Muller, Anna M. Doorn, Ismail Y. Rabbi, Wolfgang Zierer, The Cassava Source–Sink project: opportunities and challenges for crop improvement by metabolic engineering, The Plant Journal, 10.1111/tpj.14865, 103 , 5, (1655-1665), (2020).
- Zehong Ding, Lili Fu, Weiwei Tie, Yan Yan, Chunlai Wu, Jing Dai, Jiaming Zhang, Wei Hu, Highly dynamic, coordinated, and stage-specific profiles are revealed by a multi-omics integrative analysis during tuberous root development in cassava, Journal of Experimental Botany, 10.1093/jxb/eraa369, 71 , 22, (7003-7017), (2020).
- Cees Bruggink, Detlef Jensen, Combining ion chromatography with mass spectrometry and inductively coupled plasma‐mass spectrometry: Annual review 2020, Analytical Science Advances, 10.1002/ansa.202000120, 2 , 3-4, (238-249), (2020).

