Real time-quaking induced conversion assay (RT-QUIC)
Arpine Sokratian, andrew.west west
Abstract
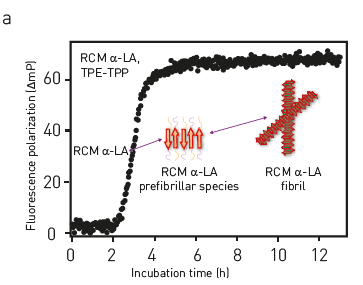
Seeded-amplification assay (SAA) method for the detection of pathology-associated aggregation-templating competent forms of alpha-synuclein and other interacting factors.
Attachments
Steps
Plate preparation
Thaw down a-syn monomer and sonicated fibril aliquots On ice, do not generate bubbles by pipetting
Measure monomer concentration via Nanodrop
Add 3µL of 10x diluted aliquot in PBS onto nanodrop piedestal;
Parameters: other proteins; coefficient extinction: 5.98; MW: 14.4 kDA
Perform two measurements and confirm <10% standard error between two measurements
If necessary, prepare 20X and 30X dilutions to confirm findings.
Equipment
| Value | Label |
|---|---|
| NanoDrop™ One/OneC Microvolume UV-Vis Spectrophotometer | NAME |
| UV-Vis Spectrophotometer | TYPE |
| Thermo Scientific | BRAND |
| ND-ONE-W | SKU |
| https://www.thermofisher.com/us/en/home.html | LINK |
Calculate the volume of monomer need following: 0.3mg/mL monomer in a reaction mix X 30µLper well
_30 ul (reaction volume) * 3 (replicates) * ____ (different conditions) = ____ul;
Dilute the monomer preparation with 2.5mg/mL in order to filter the protein preparation for a high efficiency
Use
Meantime measure concentration of sonicated fibrils: prepare serial dilutions: 2x; 4x; 8x in3Molarity (M) Guanidine HCL, volume: 10µL
Add 3µL onto nanodrop piedestal;
Parameters: other proteins; coefficient extinction: 5.98; MW: 14.4 kDA
Blank against 3Molarity (M) Guanidine HCL;
Perform two measurements for each dilution and confirm <10% standard error between two measurements
Equipment
| Value | Label |
|---|---|
| NanoDrop™ One UV-Vis Spectrophotometer | NAME |
| spectrophotometer | TYPE |
| Thermo Scientific | BRAND |
| ND-ONE-W | SKU |
| Sample Volume (Metric): Minimum 1µL; Spectral Bandwidth: ≤1.8 nm (FWHM at Hg 254 nm); System Requirements: Windows™ 8.1 and 10, 64 bit; Voltage: 12 V (DC); Wavelength Range: 190–850 nm | SPECIFICATIONS |
Measure DLS data for sonicated fibrils
Place single zirconia bead into each well of the
Prepare serial dilutions of sonicated fibrils:
Prepare solution: 1mg/mLmeasured PFFs (20µL volume);
-
10 ug/ml: 10 ul PFFs + 990 ul of PBS;
-
1000 ng/ml: 10 ul PFFs + 90 ul of PBS;
-
100 ng/ml: 10 ul PFFs + 90 ul of PBS;
-
10 ng/ml: 10 ul PFFs + 90 ul of PBS;
-
1 ng/ml: 10 ul PFFs + 90 ul of PBS;
-
100 pg/ml: 10 ul PFFs + 90 ul of PBS;
-
10 pg/ml: 10 ul PFFs + 90 ul of PBS;
-
1 pg/ml: 10 ul PFFs + 90 ul of PBS;
-
100 fg/ml: 10 ul PFFs + 90 ul of PBS;
-
10 fg/ml: 10 ul PFFs + 90 ul of PBS;
Prepare reaction mix:
0.3mg/mL monomer in a reaction mix + 10micromolar (µM) 30µL reaction * triplicates * sample number (include standard curve of PFFs serial dilutions, 10 reactions in triplicates)
[total volume * 0.1% = total volume - (total volume * 0.1%)] - [volume of filtered monomer at __ mg/mL after filter] - [ThT volume] = __ volume of PBS
Prepare a standard curve using PFFs dilutions from
-
100 ng/ml: 10 ul of #2 (from step 8) + 90 ul of reaction mix;
-
10 ng/ml: 10 ul of #3 (from step 8) + 90 ul of reaction mix;
-
1 ng/ml: 10 ul of #4 (from step 8) + 90 ul of reaction mix;
-
100 pg/ml: 10 ul of #5 (from step 8) + 90 ul of reaction mix;
-
10 pg/ml: 10 ul of #6 (from step 8) + 90 ul of reaction mix;
-
1 pg/ml: 10 ul of #7 (from step 8) + 90 ul of reaction mix;
-
100 fg/ml: 10 ul of #8 (from step 8) + 90 ul of reaction mix;
-
10 fg/ml: 10 ul of #9 (from step 8) + 90 ul of reaction mix;
-
1 fg/ml: 10 ul of #10 (from step 8) + 90 ul of reaction mix;
-
PBS: 10 ul of PBS + 90 ul of reaction mix;
Transfer reaction to the plate: start filling the plate with a standard curve samples (seal the portion with a sealing foil)
Proceed the experimental samples and cover the whole plate with a sealing foil. Spin down the plate 500x g,0h 0m 0s
for 0h 0m 30s
Equipment
| Value | Label |
|---|---|
| Eppendorf™ 022620572 | NAME |
| Microcentrifuges with a plate rotor | TYPE |
| Eppendorf | BRAND |
| 13-690-003 | SKU |
| https://www.fishersci.com/us/en/home.html | LINK |
Example of the plate layout:
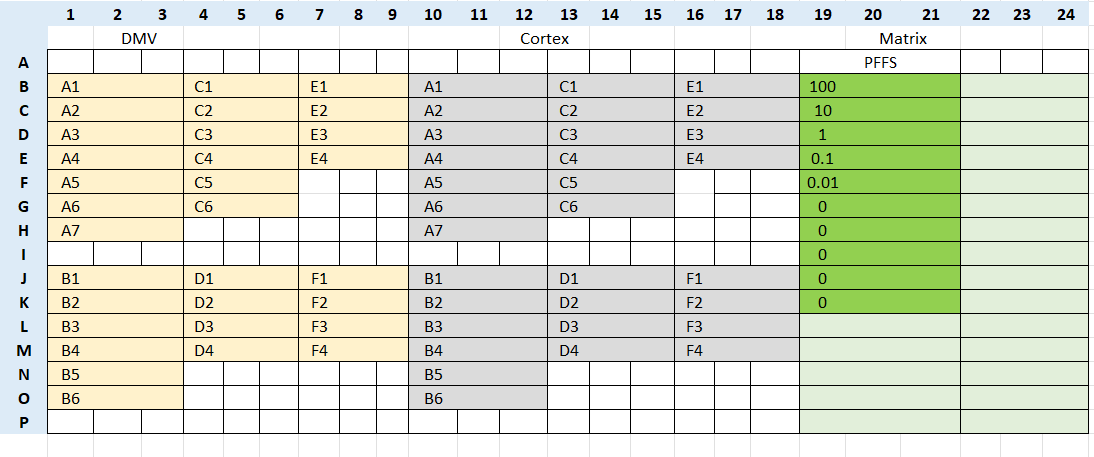
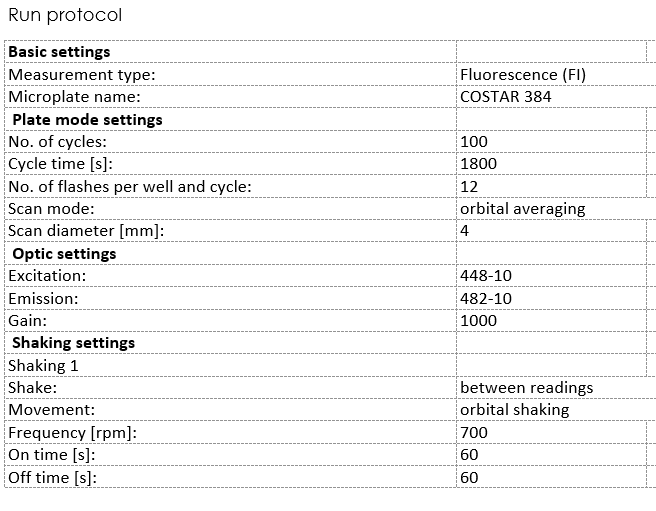
Set up the program on a plate reader
Equipment
| Value | Label |
|---|---|
| Omega - RT-QuIC / PRION Version - Fluorescence Base | NAME |
| BMG Fluostar plate reader | TYPE |
| BMG | BRAND |
| 0415-102P | SKU |
| https://www.bmglabtech.com/clariostar-plus/?utm_term=clariostar&utm_campaign=usa.roi.microplatereaders+products&utm_source=adwords&utm_medium&gclid=Cj0KCQjw4ImEBhDFARIsAGOTMj9ZLSjhHBhKWAli17cxj6V54U96GSg3qsfuFrfP1cETTLgLbn-kdUIaAkovEALw_wcB | LINK |
Data analysis
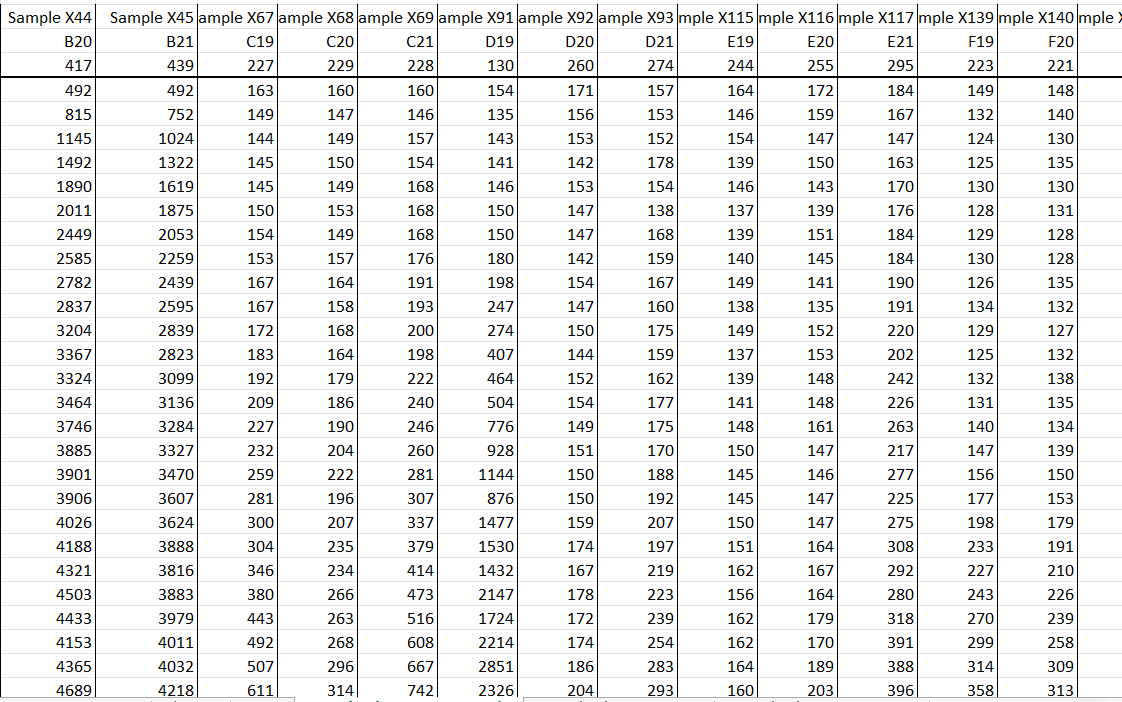
The main advantage of qQuiC analysis is capability to calculate exact amount of fibril forming units in pathological samples relatively to an appropriate standard curve. The approach of calculation FFUs consists of using standard curve with a range of serial dilutions of evaluated ssFibrils spiked in original matrix of the sample. Obtained ThT FL units for each time point and dilution are needed to define an appropriate threshold value needed to define CTvalues. To find a threshold value we built a program that can find an increase in the rates of changes greater than X present for fluorescence values in each replicate. The program looks at three points at a time and compares the rate of change from the first point to the second point against the second point to the third point. Once it finds the point where the rate of change increases by X present from one jump to the next the program stores that value to calculate the threshold value for each sample at X percent as the cutoff difference in the rates of change. The code loops through all of the replicates >30 times to check every percent threshold for X from 10% to 30% and returns the threshold value that yield the highest R-squared in a standard curve that is made from the mean CT values for each replicate. When threshold value is found CTvalue for each dilution can be determined as a time when ThT fluorescence reaches to a threshold. If R-square is higher than 0.8 than efficacy of the reaction is enough to be able to calculate FFUs. To define FFUs of experimental samples we applied an extracted equation from linear regression curve. Code for a program https://github.com/west-lab/FibrilOptimization/blob/master/FibrilPaperCode_main.py
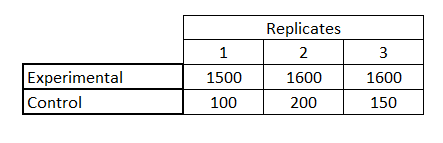
This step is required to QC the reaction by evaluating the standard curve and finding the Ct values for experimental conditions
Use mean half-time of the reaction by analyzing the Ct values of the standard curve
Use the extracted data to generate a plot or a group analysis via GraphPad