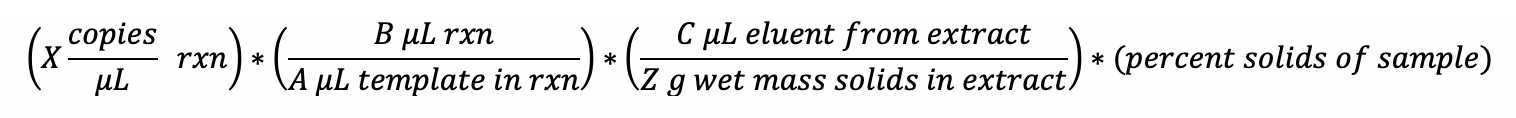High Throughput SARS-COV-2, PMMOV, and BCoV quantification in settled solids using digital RT-PCR
Krista Wigginton, Alexandria B Boehm, Aaron Topol (Verily Life Sciences), marlene.wolfe, Brad White (Verily Life Sciences)
Disclaimer
DISCLAIMER – FOR INFORMATIONAL PURPOSES ONLY; USE AT YOUR OWN RISK
The protocol content here is for informational purposes only and does not constitute legal, medical, clinical, or safety advice, or otherwise; content added to protocols.io is not peer reviewed and may not have undergone a formal approval of any kind. Information presented in this protocol should not substitute for independent professional judgment, advice, diagnosis, or treatment. Any action you take or refrain from taking using or relying upon the information presented here is strictly at your own risk. You agree that neither the Company nor any of the authors, contributors, administrators, or anyone else associated with protocols.io, can be held responsible for your use of the information contained in or linked to this protocol or any of our Sites/Apps and Services.
Abstract
This process instruction describes the steps for quantitative analysis of nucleic acid from SARS-CoV-2 with a triplex Reverse Transcriptase droplet digital Polymerase Chain Reaction (RT-ddPCR) assay targeting the N Gene, S Gene and ORF1a and a duplex assay targeting Bovine Coronavirus Vaccine (BCoV) and Pepper Mild mottle virus (PMMoV) in extracted and purified RNA samples from solid wastewater samples for population level SARS-CoV-2 community surveillance. RT-ddPCR is a modified version of conventional RT-PCR workflows which involves separating the reaction mixture into many partitions (~20,000) before thermal cycling which allows for direct absolute quantification of the target RNA molecules.
This protocol uses RNA extracted using this protocol: High Throughput RNA Extraction and PCR Inhibitor Removal of Settled Solids for Wastewater Surveillance of SARS-CoV-2 RNA. That RNA is generated from samples subjected to pre-analytical steps outlined in: High Throughput pre-analytical processing of wastewater settled solids for SARS-CoV-2 RNA analyses.
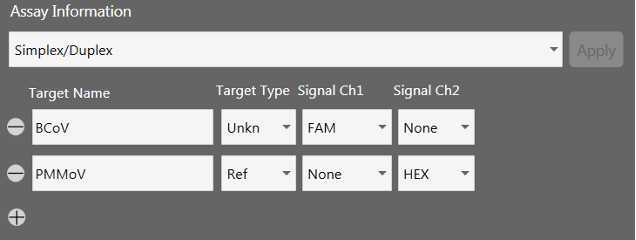
This protocol describes 2 separate PCR reactions, one containing primer/probe mixtures targeting the three SARS-CoV-2 targets and one containing primer/probe mixtures targeting BCoV and PMMoV. BCoV is spiked into samples before nucleic acid extraction and serves as a process control as well as an indicator of PCR inhibition. PMMoV is an enveloped virus which is abundant in human fecal waste and serves as an endogenous control for data normalization. PMMoV RNA is abundant at such high levels in wastewater samples that the samples must be diluted by a factor of 100 before quantification. The readout of this assay is a concentration of each target in the extracted RNA samples (copies/uL).
Scope
This process instruction applies to quantitative analysis of nucleic acid from SARS-CoV-2 RNA from solid wastewater samples with ddPCR using a Bio-Rad AutoDG Droplet Digital PCR system consisting of the AutoDG Automated Droplet Generator and the QX200 droplet reader.
Steps
Preparation (both assays)
Retrieve all kit components from the One-Step RT-ddPCR advanced kit for probes from the -20°C freezer and thaw the components -20On ice.
Retrieve ddPCR positive control aliquots (50 copies per uL gRNA and 100 copies per µL BCoV and PMMoV gene blocks) from the -80°C freezer and thaw -20On ice
For re-running frozen plates only:
Thaw the RNA storage plate -20On ice. Before analysis of thawed frozen samples, ensure that the plate is adequately sealed and briefly vortex and centrifuge the plate to ensure the samples are well mixed.
Keep extracted RNA samples -20On ice or in a cold block from the freezer at all times.
ddPCR master mix preparation (SARS-CoV-2 targets)
Prepare SARS-CoV-2 Target Master Mix
Prepare a working stock of fresh nuclease free water by pouring from a 50 mL Ambion Nuclease Free water into a 5mL eppendorf tube.
Store Master Mix components -20On ice as much as possible during the preparation process.
Briefly vortex One-Step RT supermix for Probes (Yellow Tube), Reverse Transcriptase (Orange Tube), and DTT (Grey Tube) to mix contents then briefly spin with the benchtop centrifuge.
In a 2mL eppendorf tube, prepare the master mix according to the table for SARS-CoV-2 quantification. Store prepared master mix on ice.
| A | B | C |
|---|---|---|
| Reagents | Volume per Well | Volume Per Plate |
| ddPCR™ One-Step RT supermix for Probes (Yellow Tube) | 5.5 µL | 580.8 µL |
| 20x SARS-CoV-2 Primer/Probe Mix | 3.3 µL | 348.48 µL |
| Reverse Transcriptase (Orange Tube) | 2.2 µL | 232.32 µL |
| 300mM DTT (Gray Tube) | 1.1 µL | 116.16 µL |
| Nuclease Free Water | 4.4 µL | 464.64 µL |
| Total Volume | 16.5 µL | 1742.4 µL |
SARS-CoV-2 ddPCR Master Mix. Volume per plate assumes 96 well plate (with 10% excess).
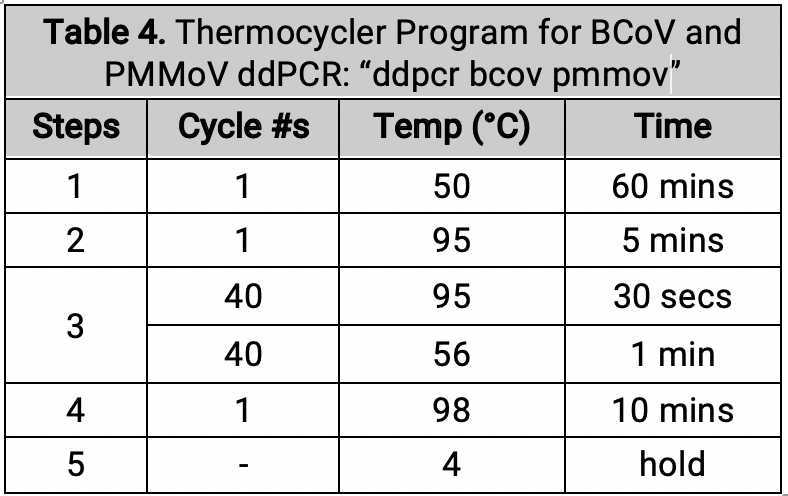
ddPCR master mix preparation (BCoV and PMMoV)
Prepare Master Mix for the BCoV/PMMoV assay
Briefly vortex One-Step RT supermix for Probes (Yellow Tube) and Reverse Transcriptase (Orange Tube) to mix contents then briefly spin with the benchtop centrifuge. Thaw DTT (grey tube).
In a 2mL eppendorf tube, prepare the master mix according to the table for PMMoV/BCoV quantification. Store prepared master mix on ice.
| A | B | C | D | E |
|---|---|---|---|---|
| Reagents | Volume per Sample | Volume Per Plate | ||
| ddPCR™ One-Step RT supermix for Probes (Yellow Tube) | 5.5 µL | 580.8 µL | ||
| BCoV/PMMoV Primer/Probe Mix | 2.2 µL | 232.32 µL | ||
| Reverse Transcriptase (Orange Tube) | 2.2 µL | 232.32 µL | ||
| 300mM DTT (Gray Tube) | 1.1 µL | 116.16 µL | ||
| Nuclease Free Water | 5.5 µL | 580.8 µL | ||
| Total Volume | 16.5 µL | 1742.4 µL |
BCoV and PMMoV ddPCR Master Mix. Volume per plate assumes 96 well plate (with 10% excess).
ddPCR plate set up (BCoV and PMMoV)
Dilute the RNA 1:100 for BCoV/PMMoV RT-ddPCR
Using a new reagent reservoir, manually pipette 198µL of nuclease free water into every well of an Eppendorf twin.tec PCR Plate labeled “100x RNA Dilution”.
Manually pipette 2µL of RNA from the stock RNA plate into each corresponding well in the dilution plate.
Seal the plate and briefly vortex the plate using a bench top plate vortexer.
Briefly spin down the plate in the Axygen Plate Centrifuge.
Store plate -20On iceuntil ready to add sample to ddPCR plate.
Transfer Master Mix and Samples to ddPCR plate (both assays)
For each assay, remove a white cold block from the freezer and place a new Bio-Rad ddPCR 96-well Plate in it.
Using a new reagent reservoir, manually pipette 16.5µL of the appropriate Master Mix to each well of each ddPCR Plate.
Transfer samples to the following wells on the plate either manually or using a liquid handling robot such as the Agilent Bravo system:
- Transfer
5.5µLof samples from columns 1-10 of the RNA plate into columns 1-10 of the ddPCR plate. - Transfer
5.5µLof the extraction controls from column 12 of the RNA plate into column 11 of the ddPCR plate. - Transfer
5.5µLof NTC (water) into A-G of column 12. - Add
5.5µLof ddPCR positive control to well H12 of the ddPCR plate (add manually even if using a liquid handler)NoteFor SARS-CoV-2 PCR: Add SARS-CoV-2 genomic RNA For PMMoV/BCoV PCR: Add PMMoV/BCoV gene block positive control
The plate layout should be:
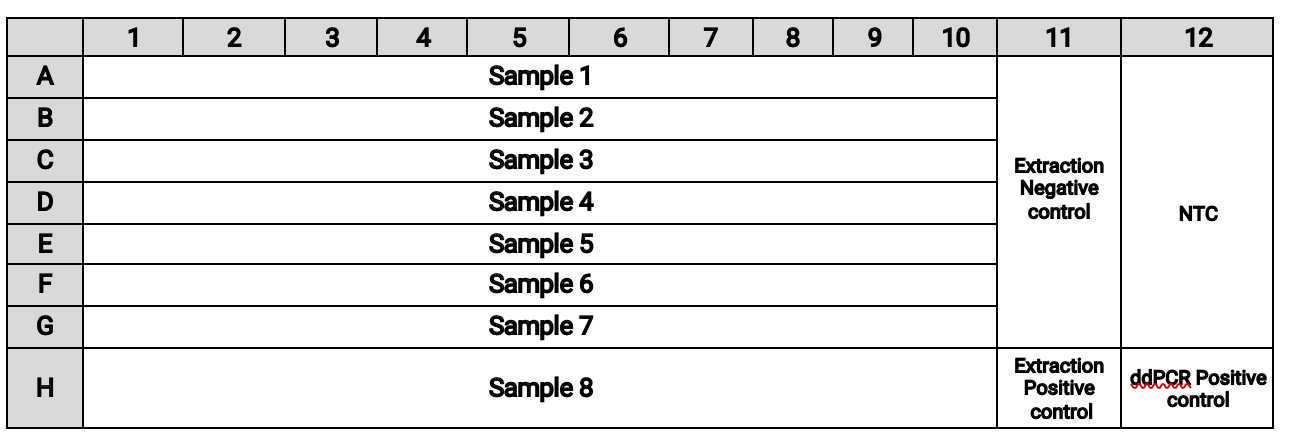
Bring the BioRad ddPCR plate with Master Mix and samples to the Bio-Rad PX1 PCR Plate Sealer and place it in the metal carrier inside the plate sealer.
Align a PCR Plate Heat Seal Foil on top of the ddPCR plate with the red line facing up.
Seal the plate by pressing the green “seal” button.
Briefly vortex the plate using a bench top plate vortexer.
Briefly spin down the plate in the bench top Axygen Plate Centrifuge.
Store the ddPCR plate on a white cold block or On ice until droplet generation.
ddPCR droplet generation (both assays)
Generate Droplets:
On the AutoDG droplet generator: select “configure plate” and then press the blue arrow in the upper left corner to highlight all columns.
Load 3 DG32 cartridge plates and 2 tip boxes with the lid removed onto the AutoDG deck. The icons on the AutoDG display will change from yellow to red if the plates and tips are oriented correctly and to green if placed correctly.
Place the sealed, vortexed, and centrifuged ddPCR plate in the Sample Plate position on the AutoDG deck.
Remove the AutoDG cooling block from the freezer and place it in the Droplet Plate slot on the AutoDG deck.
For the Droplet Plate: label a new Bio-Rad ddPCR™ 96-Well Plate and place it in the black cooling block.
Click “Generate Droplets” to begin droplet generation. Droplet generation will take approximately 0h 45m 0s.
When droplet generation is done, open the lid and remove the droplet plate from the black cold block and place it in the metal carrier inside the plate sealer.
Align a PCR Plate Heat Seal Foil on top of the droplet plate with the red line facing up.
<img src="https://static.yanyin.tech/literature_test/protocol_io_true/protocols.io.b2mjqc4n/f39ybe9nf2.jpg" alt="Photograph showing proper setup of samples and consumables in the AutoDG. After droplet generation, samples have been emulsified and deposited into the "Droplet Plate," which is ready for thermocycling." loading="lazy" title="Photograph showing proper setup of samples and consumables in the AutoDG. After droplet generation, samples have been emulsified and deposited into the "Droplet Plate," which is ready for thermocycling."/>
Seal the plate by pressing the green “seal” button.
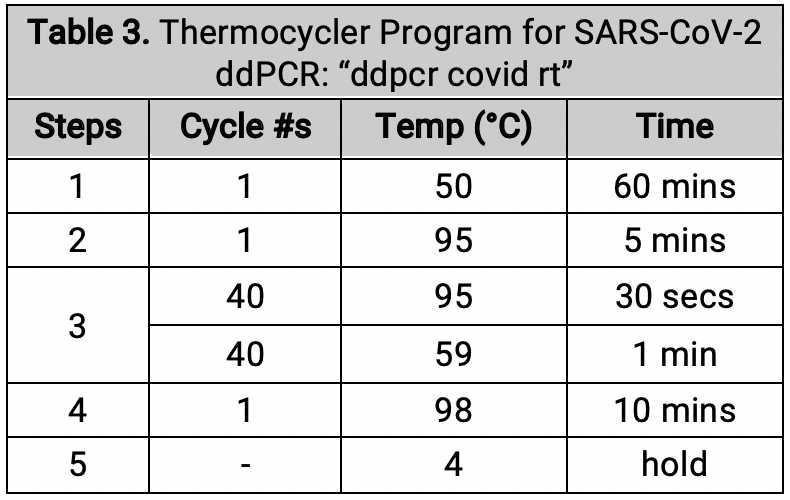
Thermocycling (SARS-CoV-2)
After the PCR program is done, store the ddPCR plate at 4°C (in the thermal cycler, on ice or in the fridge) until droplet reading.
Sample storage
After the PCR program is done, store the ddPCR plate at 4°C (in the thermal cycler, on ice or in the fridge) until droplet reading.
Read Droplets (both assays)
Before running the plate on the QX200 droplet reader, open the drawer on the left side of the instrument and check that there is adequate droplet reader oil and that the waste bottle is not overly full.
Load template from previous plate of the same assay - ensure that all parameters (dye, absorbance etc.) are correct for the assay used and that sample names are assigned to the appropriate wells. Save as with a new plate name.
Name the file with a name clearly describing which plate you are reading.
Remove the plate from the thermal cycler and secure it in the droplet reader by placing it in the stage, placing the metal brace on top of it and pressing down the black plastic tabs on either side of the brace.
Click “Run” in Quantasoft to commence droplet reading.
Select “Ok” when the next dialog box comes up. Do not change the settings from “Columns” and “FAM/HEX”.
SARS-CoV-2 Post-Processing Analysis
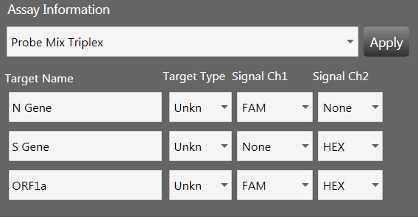
SARS-CoV-2 triplex assay analysis
Open QuantaSoft™ Analysis Pro Software
Click on the Table Menu Button on the right side of the Well Data table and select Export to CSV to export data.
Save the QuantaSoft Analysis
Open the .qlp file associated with the run
At this point, if sample names were not added at the start of the run, fill in the sample name by selecting the wells, typing in sample ID and click “Apply”.
If wells need to be deselected: Go to the Plate Editor tab, select wells to exclude and click “Clear Selected Well”.
Go through every well to ensure that:
- The droplets are designated properly per well
- There are >10,000 droplets per well
If any well does not have the appropriate droplets, go to the Plate Editor tab, select the affected well and click “Clear Selected Well”.
On the 2D Amplitude tab, select Merged Wells on the left side of the page
BCoV/PMMoV post-processing analysis
For BCoV and PMMoV analysis:
Open QuantaSoft™ Analysis Pro Software
Open the .qlp file associated with the run
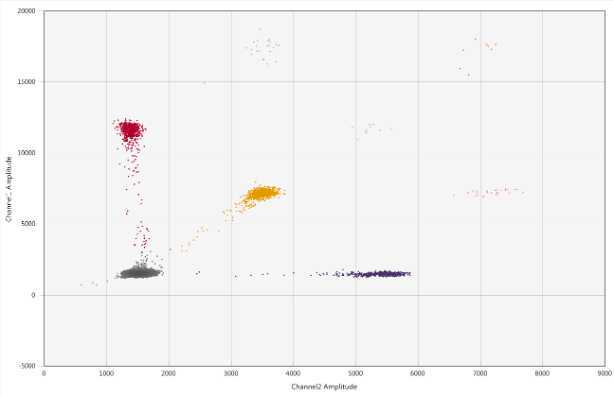
.On the 1D Amplitude tab, make sure that all wells are highlighted and select the Threshold Multiple Wells button to automatically set the threshold between the positive and negative populations
BCoV threshold = 5000
PMMoV threshold = 1700
Go through every well to ensure that:
-
The threshold accurately differentiates positive and negative droplets
-
There are >10,000 droplets per well
If any well does not have the appropriate droplets, go to the Plate Editor tab, select the affected well and click “Clear Selected Well”.
On the 1D Amplitude tab, select Merged Wells on the left side of the page
Click on the Table Menu Button on the right side of the Well Data table and select Export to CSV to export data. Save the CSV file.
Save the QuantSoft Analysis
Dimensional Analysis and Quality Control
For dimensional analysis to express the results of each assay in terms of gene copies/dry weight solids:
Begin with the concentration provided by the QuantaSoft software and reported in the CSV, as expressed in gc/uL of reaction.
To determine the concentration in cp/g dry weight: