Chronic intermittent hypoxia remodels catecholaminergic innervation in mouse atria
Ariege Bizanti, Yuanyuan Zhang, Kohlton Bendowski, Jin Chen, Maci Heal, Anas Mistareehi, Zulema Toledo, Scott W. Harden, David Gozal, Richard Christie, Peter J. Hunter, Julian F.R. Paton, Zixi Jack Cheng
Abstract
Chronic intermittent hypoxia (CIH, a model for sleep apnea) is a major risk factor for several cardiovascular diseases (CVD). Autonomic imbalance (sympathetic overactivity and parasympathetic withdrawal) has emerged as a causal contributor of CIH-induced CVD. Previously, we showed that CIH remodels the parasympathetic pathway. However, whether CIH induces remodeling of the cardiac sympathetic innervation remains unknown. Mice (male, C57BL/6J, 2-3 months) were exposed to either room air (RA, 21% O2) or CIH (alternating 21% and 5.7% O2, every 6 min, 10h/day) for 8-10 weeks. Flat-mounts of their left and right atria were immunohistochemically labeled for tyrosine hydroxylase (TH, a sympathetic marker). Using a confocal microscope (or a Zeiss M2 Imager) and a Neurolucida 360 digitization and tracing system, we scanned both the left and right atria and quantitatively analyzed the sympathetic axon density in both groups. The segmentation data was mapped onto a 3D mouse heart scaffold using the NIH SPARC Scaffold Mapping tool. Our findings indicated that CIH significantly remodeled the TH-IR innervation of the atria by increasing its density at the sinoatrial node, the auricles, and the major veins attached to the atria (p <0.05, n=7). Additionally, CIH increased the branching points of TH-IR axons while decreasing the distance between varicosities. Following CIH, abnormal patterns of TH-IR axons around intrinsic cardiac ganglia were also found. We postulate that the increased sympathetic innervation may further amplify the effects of enhanced CIH-induced central sympathetic drive to the heart. Our work provides an anatomical foundation for the understanding of CIH-induced autonomic imbalance.
The data generated with this protocol will be available on the SPARC portal at https://doi.org/10.26275/qdv4-0vx5
Steps
Hypoxia exposure
Mice were placed in cages (2 per cage) inside identical chambers with oxygen levels controlled by computerized environmental chambers (30 × 20 × 20 inches; Oxycycler A44XO, BioSpherix, Redfield, NY) for automatic delivery of intermittent hypoxia.
The light and dark cycle of the room was set to 12:12h (light 7:00 AM to 7:00 PM) at 21-22o C. CIH consisted of alternating 21% and 5.7% O2, every 6 min during the 10-hour light cycle while animals were maintained at 21% O2 for the rest of the circadian cycle for ~8-10 weeks.
Ambient CO2 in the chambers was periodically monitored and maintained at 0.03% by adjusting overall chamber ventilation.
Humidity was measured and maintained at 40-50%. Control animals were placed in the same room and exposed to continuous circulating room air (O221%) in another chamber.
Heart dissection and flat-mount preparation
Mice were first deeply anesthetized with isoflurane (5%; 5-10 minutes), with an oxygen flow rate of 1 liter per minute. When the animals were not responsive to the hind-toe pinch withdrawal reflex, the chest was opened and an injection of heparin (100 units) was made in the apex of the left ventricle. After the inferior vena cava was cut to drain the blood and perfusate, a needle was inserted into the left ventricle and the perfusion began. The mice were first perfused with 300 ml of 37-40 °C phosphate-buffered saline (0.1 M PBS, pH = 7.4) followed by 150 ml paraformaldehyde (4%; 4 °C).
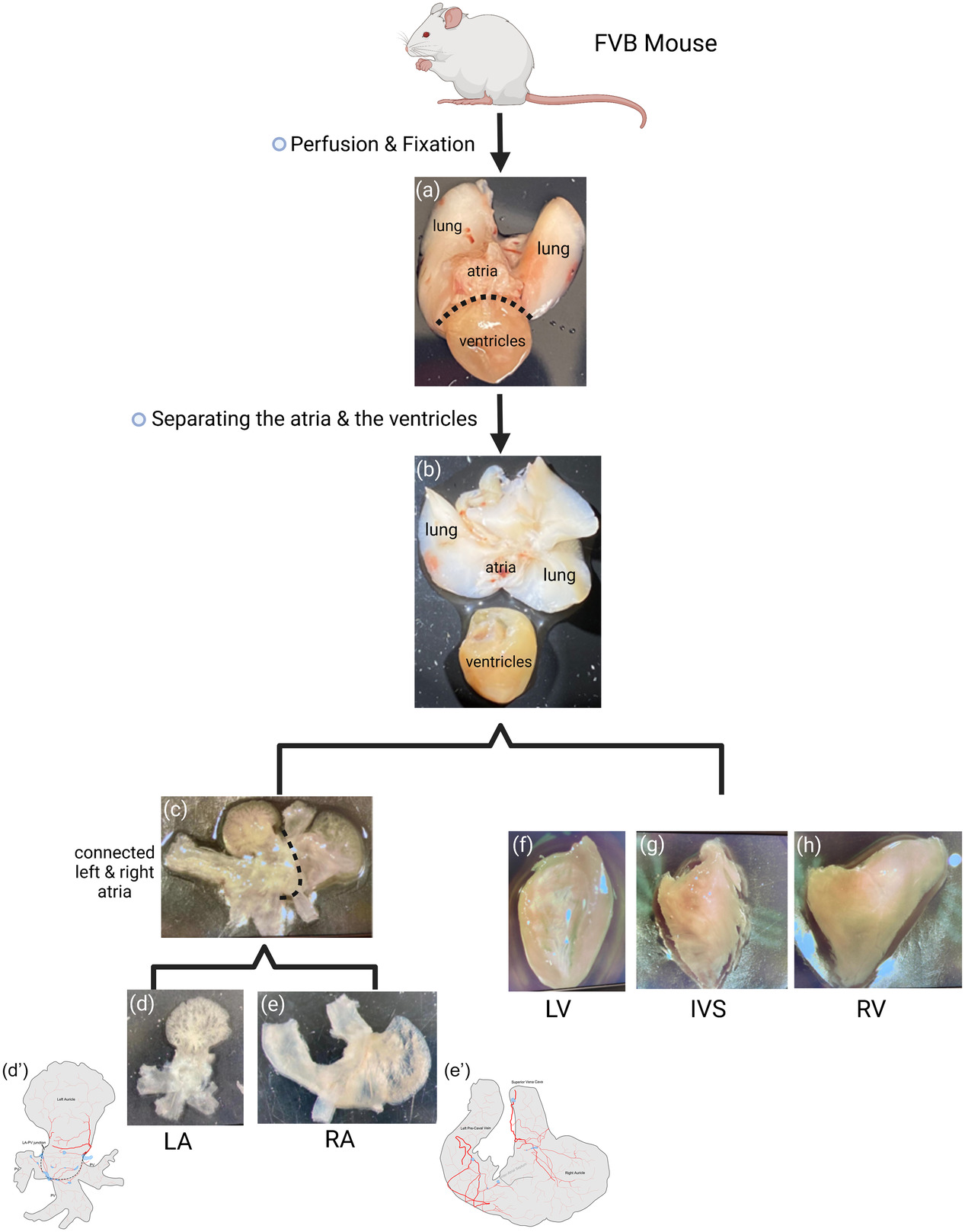
The tissues were separated in the same manner that were performed as described previously (1) or as shown in Figure 1 of (2).
The heart, lungs, and trachea were removed immediately after perfusion and stored in 4 % paraformaldehyde for at least 24 hours at 4 ºC. The atria and ventricles were separated at the atrial-ventricular groove, and the left and right atria and ventricles were separated. The left atrium was prepared with the entrance portions of all pulmonary veins (PVs) attached and the right atrium was prepared with the superior vena cava (SVC), inferior vena cava (IVC) and left precaval vein (LPCV) attached.
Immunohistochemistry (IHC)
Dissected tissues were washed 3 times for 15 min in 0.01 M PBS (pH = 7.4) in a 24-well plate on an orbital shaker to remove the remaining fixative.
To prevent non-specific binding and enhance antibody penetration, the atria were fully submerged in a blocking solution (2% bovine serum albumin, 10% normal donkey serum, 2% Triton X-100, 0.08% NaN3 (in 0.1 M PBS, pH = 7.4)) for 2-3 days at 4 ºC.
Anti-TH primary antibodies (12 μl/ml) were added to primary solution (2% bovine serum albumin, 4% normal donkey serum, 0.08% Triton X-100, .008% NaN3in 0.1 M PBS, pH=7.4) and incubated for 48 h.
Tissues were thoroughly washed 6 times for 10 min in PBST (phosphate buffered saline with Triton X-100; 0.5% Triton X-100 in 0.01 M PBS) to remove unbound primary antibodies.
Tissues were then kept in the dark and incubated in a fluorescent secondary antibody solution (Alexa Fluor 488; Invitrogen, Waltham, MA, Cat# A21206, RRID:AB_2535792; 1:50) overnight at room temperature. Unbound secondary antibodies were removed by washing 6 times for 10 minutes in PBS at room temperature.
Tissues were then mounted onto glass slides and flattened with lead weights (2.5 kg x4) for 2 day.
Slides were dehydrated in four ascending concentrations of ethanol (75%, 95%, 100%, and 100%) for 2 min in each concentration, followed by 2 × 10 min washes in 100% xylene.
Coverslips were then attached using DEPEX mounting medium (Electron Microscopy Sciences #13514) and allowed to dry overnight.
| A | B | C | D | E | F |
|---|---|---|---|---|---|
| Antibody | Concentration | Host | Company | Catalog | RRID |
| Anti-TH | 1:100 12 μL/mL | Rabbit | Pel-Freeze | Cat# P40101-0 | AB_461064 |
| Anti-TH | 1:100 12 μL/mL | Sheep | Millipore | AB1542 | AB_90755 |
| Anti- PGP9.5 | 1:100 12 μL/mL | Rabbit | Abcam | ab108986 | AB_10891773 |
| Alexa Fluor 594 anti-sheep | 1:50 24 μL/mL | Donkey | Invitrogen | A-11016 | AB_2534083 |
| Alexa Fluor 488 | 1:50 24 μL/mL | Donkey | Invitrogen | A21206 | AB_2535792 |
Table 1: List of antibodies used
Image acquisition
Hundreds of overlapping maximum projection images from stacks of optical sections (z-step: 1.5 μm) from each tissue were captured using a Zeiss M2 Imager (20x lens; NA 0.8) and stitched back together seamlessly to yield full photo montages of the right and left atria and ventricles. An LED light source with a 488 nm wavelength was used to visualize the CGRP-IR axons in the tissues.
A Leica TCS SP5 laser-scanning confocal microscope (40x oil immersion lens; NA 1.25; z-step: 1 μm) was used to capture detailed images of CGRP-IR axons and their targets in selected locations of the heart. An argon–krypton laser (488 nm) was used to visualize TH-IR axons in the tissue, and a helium-neon laser (543 nm) was used to detect the autofluorescence of the tissues in the background (e.g., cardiac muscles, ganglionic cells, and blood vessels).
Modifications, including brightness and contrast adjustments, and scale bar additions, were performed using Photoshop or FIJI ImageJ software
Density quantification
To quantify the regional density of TH-IR fibers in the atria, we segregated images into specific regions of interest (ROIs) similar to our previous work (3): SAN region, AVN region (the atrial region at the interatrial septum near the AVN) , SVC, IVC, right outer and inner auricle, LA-PV junction, left PV, middle PV, right PV, left outer and inner auricle.
Six window frames 1000x1000px which included all z-steps were selected and averaged from each region of each group. The steps of the density quantification were as follows: 1) subtracted the background with radius of 80 pixels to reduce noise and enhance contrast. 2) applied particle removal to remove small debris. 3) applied a binary threshold (Otsu method) to isolate immunoreactive structures and then images were skeletonized. 4) quantified the signal above the threshold. 5) averaged the signal of different ROI windows using six counting frames. 6) divide the number of signal pixels by ROI area. 7) ran the Shapiro-Wilk normality test. S
Statistical significance of the difference between the means was performed using one-way ANOVA and Tukey’s HSD (Honestly Significant Difference). Data were expressed as mean density % +/- SD. Statistical significance was accepted at P value <0.05.
Heatmaps of axon intensity were created after applying a modified version of the freely available open-source automated software algorithm that traces and quantifies axons (Axon tracer plugin , ImageJ) (4).
Axon tracing and quantification
Neurolucida 360 tracing : We used Neurolucida 360® software (https://www.mbfbioscience.com/neurolucida360, RRID:SCR_016788) to digitize and trace TH-IR axonal structures. Our axon tracing follows the steps provided in our previous protocols: https://www.protocols.io/view/topographical-mapping-of-sympathetic-postganglioni-n92ldzbmxv5b/v2, Neurolucida Explorer®(https://www.mbfbioscience.com/neurolucida-explorer, RRID:SCR_017348) was used in combination to perform morphometric analysis including complexity (number of bifurcation points per unit of area) and varicosity distance.
Zeiss Arivis Vision 4D was utilized to autotrace large size deconvolution data of the whole right and left atria through a customized pipeline. The analysis pipeline was used for quantification of the 3D image data and complete tracing. Features selected to measure included the axons total length of axon, average tortuosity, and number of bifurcation points.
Anatomical mapping onto a 3D heart scaffold
To map the tracing data onto a 3D anatomical heart scaffold for a more intuitive 3D spatial distribution of atrial axon innervation, we utilized the open-source software framework Musculoskeletal atlas project (MAP) https://map-client.readthedocs.io/en/latest/.
We created a workflow to map the traced TH-IR axon innervation from flat mount preparations of the whole right and left atrium in both RA and CIH conditions.
The generic mouse heart scaffold was created based on a set of anatomical and mathematical parameters from the standard shape and size of mouse hearts using an open-source Python library called ScaffoldMaker created by Auckland Bioengineering Institute (https://github.com/ABI-Software/scaffoldmaker, RRID:SCR_019003).
Preparation of workflow and mapping data demonstration tutorials are available on https://docs.sparc.science/docs/scaffold-mapping-tools-mapping-image-data.
To align the scaffold into the same coordinate space as the data, fiducial annotations with similar terminologies were added to both. These annotations were specific to each tissue morphological edge.
Geometrical fitting of anatomical scaffold to digitized contours and fiducial marker points resulted in a fitted and flattened left atrium.
Scaffold Mapping: Data is mapped onto the scaffold using the calculated geometric coordinates relative to the fitted scaffold.

